Artificial Intelligence in Health Care: The Hope, the Hype, the Promise, the Peril (2022)
Chapter: 5 Artificial Intelligence Model Development and Validation
5
ARTIFICIAL INTELLIGENCE MODEL DEVELOPMENT AND VALIDATION
Hongfang Liu, Mayo Clinic; Hossein Estiri, Harvard University; Jenna Wiens, University of Michigan; Anna Goldenberg, University of Toronto; Suchi Saria, Johns Hopkins University; and Nigam Shah, Stanford University
INTRODUCTION
This chapter provides an overview of current best practices in model development and validation. It gives readers a high-level perspective of the key components of development and validation of artificial intelligence (AI) solutions in the health care domain but should not be considered exhaustive. The concepts discussed here apply to the development and validation of an algorithmic agent (or, plainly, an algorithm) that is used to diagnose, prognose, or recommend actions for preventive care as well as patient care in outpatient and inpatient settings.
Overall, Chapters 5, 6, and 7 represent the process of use case and needs assessment, conceptualization, design, development, validation, implementation, and maintenance within the framework of regulatory and legislative requirements. Each of these chapters engages readers from a different perspective within the ecosystem of AI tool development, with specific considerations from each lens (see Figure 1-6 in Chapter 1).
A complete evaluation of an AI solution in health care requires an assessment of utility, feasibility given available data, implementation costs, deployment challenges, clinical uptake, and maintenance over time. This chapter focuses on the process necessary to develop and validate a model, and Chapter 6 covers the issues of implementation, clinical use, and maintenance.
This chapter focuses on how to identify tasks that can be completed or augmented by AI, on challenges in developing and validating AI models for health care, and on how to develop AI models that can be deployed. First, all stakeholders must understand the needs coming from clinical practice, so that proposed AI
systems address the needs of health care delivery. Second, it is necessary that such models be developed and validated through a team effort involving AI experts and health care providers. Throughout the process, it is important to be mindful of the fact that the datasets used to train AI are heterogeneous, complex, and nuanced in ways that are often subtle and institution specific. This affects how AI tools are monitored for safety and reliability, and how they are adapted for different locations and over time. Third, before deployment at the point of care, AI systems should be rigorously evaluated to ensure their competency and safety, in a process similar to that done for drugs, medical devices, and other interventions.
Illustrative Use Case
Consider the case of Vera, a 60-year-old woman of Asian descent with a history of hypertension, osteoporosis, diabetes, and chronic obstructive pulmonary disease, entering the doctor’s office with symptoms of shortness of breath and palpitations. Her primary care physician must diagnose and treat the acute illness, but also manage the risks associated with her chronic diseases. Ideally, decisions regarding if to treat are guided by risk stratification tools, and decisions of how to treat are guided by evidence-based guidelines. However, such recommendations are frequently applied to patients not represented in the data used in those recommendations’ development. For example, best practices to manage blood pressure come from randomized controlled trials enrolling highly homogeneous populations (Cushman et al., 2016), and methods for risk stratification (e.g., assessment via the atherosclerotic cardiovascular disease risk equation) frequently do not generalize well to diverse populations (Yadlowsky et al., 2018). Moreover, there are care delivery gaps from clinical inertia, provider familiarity with treatments, and patient preferences that result in the lack of management or under-treatment of many conditions.
Vera’s experience would improve if an automated algorithm could support health care teams to accurately (1) classify and precisely recognize existing conditions (accounting for her age, race, and genetic makeup more broadly), (2) predict risks of specific events (e.g., 1-year risk of stroke) given the broad context (e.g., comorbidities such as osteoporosis) and not just the current symptoms, and (3) recommend specific interventions.
Even for well-understood clinical situations, such as the monitoring of surgical site infections, current care delivery falls woefully short. For example, the use of automated text-processing and subsequent reasoning on both the text-derived content as well as other structured data in the electronic health record (EHR) resulted in better tools for monitoring surgical site infections in real time (Ke et al., 2017; Sohn et al., 2017). Such automation can significantly increase the capacity
TABLE 5-1 | Example of Artificial Intelligence Applications by the Primary Task and Main Stakeholder
| Classify (Diagnose) | Predict (Prognose) | Treat | |
|---|---|---|---|
| Payers | Identify which patients will not adhere to a treatment plan | Estimate risk of a “no-show” for a magnetic resonance imaging appointment | Select the best second line agent for managing diabetes after Metformin given a specific clinical history |
| Patients and caregivers | Estimate risk of having an undiagnosed genetic condition (e.g., familial hypercholesterolemia) | Estimate risk of a postsurgical complication | Identify a combination of anticancer drugs that will work for a specific tumor type |
| Providers | Identify patients with unrecognized mental health needs | Determine risk of acute deterioration needing heightened care | Establish how to manage an incidentally found atrial septal defect |
of a care team to provide quality care or, at the very least, free up their time from the busy work of reporting outcomes. Below, additional examples are discussed regarding applications driving the use of algorithms to classify, predict, and treat, which can guide users in figuring out for whom to take action and when. These applications may be driven by needs of providers, payers, or patients and their caregivers (see Table 5-1).
MODEL DEVELOPMENT
Establishing Utility
One of the most critical components of developing AI for health care is to define and characterize the problem to be addressed and then evaluate whether it can be solved (or is worth solving) using AI and machine learning. Doing so requires an assessment of utility, feasibility given available data, implementation costs, deployment challenges, clinical uptake, and maintenance over time. This chapter focuses on the process necessary to develop and validate a model, and Chapter 6 covers the issues of implementation, clinical use, and maintenance.
It is useful to think in terms of how one would act given a model’s output when considering the utility of AI in health care. Factors affecting the clinical utility of a predictive model may include lead time offered by the prediction, the existence of a mitigating action, the cost and ease of intervening, the logistics of the intervention, and incentives (Amarasingham et al., 2014; Meskó et al., 2018; Yu et al., 2018). While model evaluation typically focuses on metrics such as positive predictive value, sensitivity (or recall), specificity, and calibration,
constraints on the action triggered by the model’s output (e.g., continuous rhythm monitoring might be constrained by availability of Holter monitors) often can have a much larger influence in determining model utility (Moons et al., 2012).
For example, if Vera was suspected of having atrial fibrillation based on a personalized risk estimate (Kwong et al., 2017), the execution of follow-up action (such as rhythm monitoring for 24 hours) depends on availability of the right equipment. In the absence of the ability to follow up, having a personalized estimate of having undiagnosed atrial fibrillation does not improve Vera’s care.
Therefore, a framework for assessing the utility of a prediction-action pair resulting from an AI solution is necessary. During this assessment process, there are several key conceptual questions that must be answered (see Box 5-1). Quantitative answers to these questions can drive analyses for optimizing the desired outcomes, adjusting components of the expected utility formulation and fixing variables that are difficult to modify (e.g., the cost of an action) to derive the bounds of optimal utility.
For effective development and validation of AI/machine learning applications in health care, one needs to carefully formulate the problem to be solved, taking into consideration the properties of the algorithm (e.g., positive predictive value) and the properties of the resulting action (e.g., effectiveness), as well as the constraints on the action (e.g., costs, capacity), given the clinical and psychosocial environment.
If Vera’s diagnosis was confirmed and subsequently the CHADS2 risk score indicated a high 1-year risk of ischemic stroke, the utility of treating using anticoagulants has to be determined in the light of the positive predictive value of the CHADS2 score and the known (or estimated) effectiveness of anticoagulation in preventing the stroke as well as the increased risk of bleeding incurred from anticoagulant use in the presence of hypertension.
Choices Before Beginning Model Development
After the potential utility has been established, there are some key choices that must be made prior to actual model development. Both model developers and model users are needed at this stage in order to maximize the chances of succeeding in model development because many modeling choices are dependent on the context of use of the model. Although clinical validity is discussed in Chapter 6, we note here that the need for external validity depends on what one wishes to do with the model, the degree of agency ascribed to the model, and the nature of the action triggered by the model.
The Two Cultures of Data Modeling and Algorithmic Modeling
Expert-driven systems attempt to capture knowledge and derive decisions through explicit representation of expert knowledge. These types of systems make use of established biomedical knowledge, conventions, and relationships. The earliest AI systems in health care were all expert-driven systems, given the paucity of datasets to learn from (e.g., MYCIN) (Shortliffe, 1974).
Modern AI systems often do not attempt to encode prior medical knowledge directly into the algorithms but attempt to uncover these relationships during model learning. Inexpensive data storage, fast processors, and advancements in machine learning techniques have made it feasible to use large datasets, which can uncover previously unknown relationships. They have also been shown to outperform expert-driven approaches in many cases, as illustrated through the latest digital medicine focus in Nature Medicine (Esteva et al., 2019; Gottesman et al., 2019; He et al., 2019). However, learning directly from the data carries the risk of obtaining models with decreased interpretability, getting potentially counterintuitive (to a clinical audience) models and models that are overfitted and fail to generalize, and propagating biased decision making by codifying the existing biases in the medical system (Char et al., 2018; Chen et al., 2019; Vellido, 2019; Wang et al., 2018).
In addition, when a model is learning from (or being fitted to) the available data, there are often conflicting views that lie at two ends of a spectrum (Breiman, 2001). At one end are approaches that posit a causal diagram of which variables influence other variables, in such a way that this “model” is able to faithfully represent the underlying process that led to the observed data—that is, we attempt to model the process that produced the data (Breiman, 2001). At the other end of the spectrum are approaches that treat the true mechanisms that produced the observed data as unknown and focus on learning a mathematical function that maps the given set of input variables to the observed outcomes. Such models merely capture associations and make no claim regarding causality (Cleophas et al., 2013).
As an example, consider the situation of taking a dataset of observed characteristics on 1 million individuals and building a model to assess the risk of a heart attack in 1 year. The causal approach would require that the variables we collect—such as age, race, gender, lipid levels, and blood pressure—have some underlying biological role in the mechanisms that lead to heart attacks. Once a model learns from the data, the variables that are most associated with the outcome “have a role” in causing the outcome; modifying those variables (e.g., lowering lipid levels) reduces the risk of the outcome (the heart attack). Typically, such models also have external validity in the sense that the risk equation learned once (e.g., from the Framingham Heart Study in New Hampshire) works in a different geography (e.g., California) decades later (Fox, 2010; Mason et al., 2012). If one is interested purely in associations, one might include every variable available, such as hair color, and also tolerate the fact that some key items such as lipid levels are missing. In this case the model—the mathematical function mapping inputs to outcomes—learns to weigh “hair color” as an important factor for heart attack risk and learns that people with white or gray hair have a higher risk of heart attacks. Clearly, this is a cartoon example, as coloring people’s hair black does nothing to reduce their heart attack risk. However, if one needs to estimate financial risk in insuring our hypothetical population of 1 million individuals, and a large fraction of that population has white or gray hair, a model need not be “causal.” As Judea Pearl writes, “Good predictions need not have good explanations. The owl can be a good hunter without understanding why the rat always goes from point A to point B” (Pearl and Mackenzie, 2018).
Ideally, one can combine prior medical knowledge of causal relationships with learning from data in order to develop an accurate model (Schulam and Saria, 2017; Subbaswamy et al., 2018). Such combination of prior data can take multiple forms. For example, one can encode the structure fully and learn the parameters from data (Sachs et al., 2005). Alternatively, one could develop models that faithfully represent what is known about the disease and learn the remaining structure and parameters (Schulam and Saria, 2017; Subbaswamy et al., 2018).
Model learning that incorporates prior knowledge automatically satisfies construct validity and is less likely to generate counterintuitive predictions. Many view the use of prior knowledge or causal relationships as impractical for any sufficiently complex problem, in part because the full set of causal relationships is rarely known. However, recent works show that even when we have established only partial understanding of causal relationships, we can improve generalizability across environments by removing spurious relationships that are less likely to be generalizable, thereby reducing the risk of catastrophic prediction failures (Schulam and Saria, 2017; Subbaswamy et al., 2018).
The topic of interpretability deserves special discussion because of ongoing debates around interpretability, or the lack of it (Licitra et al., 2017; Lipton, 2016; Voosen, 2017). Interpretability can mean different things to the individuals who build the models and the individuals who consume the model’s output. To the model builder, interpretability often means the ability to explain which variables and their combinations, in what manner, led to the output produced by the model (Friedler et al., 2019). To the clinical user, interpretability could mean one of two things: a sufficient enough understanding of what is going on, so that they can trust the output and/or be able to get liability insurance for its recommendations; or enough causality in the model structure to provide hints as to what mitigating action to take. The Framingham risk equation satisfies all three meanings of interpretability (Greenland et al., 2001). Interpretability is not always necessary, nor is it sufficient for utility. Given two models of equal performance, one a black box model and one an interpretable model, most users prefer the interpretable model (Lipton, 2016). However, in many practical scenarios, models that may not be as easily interpreted can lead to better performance. Gaining users’ trust has often been cited as a reason for interpretability (Abdul, 2018). The level of users’ trust highly depends on the users’ understanding of the target problems. For example, one does not need to have an interpretable model for a rain forecast in order to rely on it when deciding whether to carry an umbrella, as long as the model is correct enough, often enough, because a rain forecast is a complex problem beyond the understanding of individuals. For non-expert users, as long as the model is well validated, the degree of interpretability often does not affect user trust in the model, whereas expert users tend to demand a higher degree of model interpretability (Poursabzi-Sangdeh, 2018).
To avoid wasted effort, it is important to understand what kind of interpretability is needed in a particular application. For example, consider the prediction of 24-hour mortality using a deep neural network (Rajkomar et al., 2018). It can take considerably more effort to train and interpret deep neural networks, while linear models, which are arguably more interpretable, may yield sufficient performance. If interpretability is deemed necessary, it is important to understand why interpretability is needed. In Rajkomar et al. (2018), the neural networks for predicting high 24-hour mortality risk were interrogated to interpret their predictions and highlight the reasons for the high mortality risk. For a patient with high 24-hour mortality risk, in one case, the reasons provided were the presence of metastatic breast cancer with malignant pleural effusions and empyema. Such engineering interpretability, while certainly valid, does not provide suggestions for what to do in response to the high-mortality prediction.
A black box model may suffice if the output was trusted, and the recommended intervention was known to affect the outcome. Trust in the output can be obtained
by rigorous testing and prospective assessment of how often the model’s predictions are correct and calibrated, and for assessing the impact of the interventions on the outcome. At the same time, prospective assessment can be costly. Thus, doing all one can to vet the model in advance (e.g., by inspecting learned relationships) is imperative.
Augmentation Versus Automation
In health care, physicians are accustomed to augmentations. For example, a doctor is supported by a team of intelligent agents, including specialists, nurses, physician assistants, pharmacists, social workers, case managers, and other health care professionals (Meskó et al., 2018). While much of the popular discussion of AI focuses on how AI tools will replace human workers, in the foreseeable future, AI will function in an augmenting role, adding to the capabilities of the technology’s human partners. As the volume of data and information available for patient care grows exponentially, AI tools will naturally become part of such a care team. They will provide task-specific expertise in the data and information space, augmenting the capabilities of the physician and the entire team, making their jobs easier and more effective, and ultimately improving patient care (Herasevich et al., 2018; Wu, 2019).
There has been considerable ongoing discussion about the level of autonomy that AI can or should have within the health care environment (Verghese et al., 2018). The required performance characteristics and latitude of use of AI models is substantially different when the model is operating autonomously versus when a human is using it as an advisor and making the final decision. This axis also has significant policy, regulatory, and legislative considerations, which are discussed in Chapter 7. It is likely that progress in health care AI will proceed more rapidly when these AI systems are designed with the human in the loop, with attention to the prediction-action pairing, and with considerations of the societal, clinical, and personal contexts of use. The utility of those systems highly depends on the ability to augment human decision-making capabilities in disease prevention, diagnosis, treatment, and prognosis.
Selection of a Learning Approach
The subfield of AI focused on learning from data is known as machine learning. Machine learning can be grouped into three main approaches: (1) supervised, (2) unsupervised, and (3) reinforcement learning. Each approach can address different needs within health care.
Supervised learning focuses on learning from a collection of labeled examples. Each example (i.e., patient) is represented by input data (e.g., demographics, vital signs, laboratory results) and a label (such as being diabetic or not). The learning algorithm then seeks to learn a mapping from the inputs to the labels that can
generalize to new examples. There have been many successful applications of supervised learning to health care. For example, Rajpurkar et al. (2018) developed an AI system that classified 14 conditions in chest radiographs at a performance level comparable to that of practicing radiologists. Based on the training data, their system learned which image features were most closely associated with the different diagnoses. Such systems can also be used to train models that predict future events. For example, Rajkomar et al. (2018) used supervised learning to learn a mapping from the structured as well as textual contents of the EHR to a patient’s risk for mortality, readmission, and diagnosis with specific International Classification of Diseases, Ninth Revision, codes by discharge (Rajkomar et al., 2018). This method of training a model is popular in settings with a clear outcome and large amounts of labeled data. However, obtaining labeled data is not always straightforward. Unambiguous labels may be difficult to obtain for a number of reasons: the outcome or classification may be ambiguous, with little interclinician agreement; the labeling process may be labor intensive and costly; or labels may simply be unavailable. In many settings, there may not be a large enough dataset to confidently train a model. In such settings, weakly supervised learning can be leveraged when noisy, weak signals are available. For example, to mitigate the burden of expensive annotations, one study used weak supervision to learn a severity score for acute deterioration (Dyagilev and Saria, 2016). In another study where it was not possible to acquire gold-standard labels, weak supervision was used to learn a disease progression score for Parkinson’s disease (Zhan et al., 2018). Various other strategies, including semi-supervised learning and active learning, can be deployed to reduce the amount of labeled data needed (Zhou, 2017).
Unsupervised learning seeks to examine a collection of unlabeled examples and group them by some notion of shared commonality. Clustering is one of the common unsupervised learning tasks. Clustering algorithms are largely used for exploratory purposes and can help identify structure and substructure in the data. For example, Williams et al. (2018) clustered data pertaining to more than 10,000 pediatric intensive care unit admissions, identifying clinically relevant clusters. Unsupervised learning can also be used to stage or subtype heterogeneous disease (Doshi-Velez et al., 2014; Goyal et al., 2018; Saria and Goldenberg, 2015). Here, the difficulty lies not in obtaining the grouping—although such techniques similarly suffer from small datasets—but in evaluating it. When given a dataset and a clustering algorithm, we always get a grouping. The challenge, then, is whether the presence of the groups (i.e., clusters) or learning that a new patient is deemed a member of a certain group is actionable in the form of offering different treatment options. Most often, the ability to reproduce the same groups in another dataset is considered a sign that the groups are medically meaningful and perhaps they should be managed differently. If the fact that a new record belongs to a certain
group allows an assignment of higher (or lower) risk of specific outcomes, that is considered a sign that the learned groups have meaning. For example, Shah et al. (2015) analyzed a group of roughly 450 patients who had heart failure with preserved ejection fraction in order to find three subgroups (Shah et al., 2015). In data that were not used to learn the groups, application of the grouping scheme sorted patients into high, medium, and low risk of subsequent mortality.
Reinforcement learning differs from supervised and unsupervised learning, because the algorithm learns through interacting with its environments rather than through observational data alone. Such techniques have had recent successes in game settings (Hutson, 2017). In games, an agent begins in some initial stage and then takes actions affecting the environment (i.e., transitioning to a new state) and receiving a reward. This framework mimics how clinicians may interact with their environment, adjusting medication or therapy based on observed effects. Reinforcement learning is most applicable in settings involving sequential decision making where the reward may be delayed (i.e., not received for several time steps). Although most applications consider online settings, recent work in health care has applied reinforcement learning in an offline setting using observational data (Komorowski et al., 2018). Reinforcement learning holds promise, although its current applications suffer from issues of confounding and lack of actionability (Saria, 2018).
LEARNING A MODEL
To illustrate the process of learning a model, we focus on a supervised learning task for risk stratification in health care. Assume we have n patients. Each patient is represented by a d-dimensional feature vector that lies in some feature space X (see rows in Figure 5-1). In addition, each patient has some label, y, representing that patient’s outcome or condition (such as being diabetic or not). In some settings, we may have only a single label for each patient; in others we may have multiple labels that vary over time. We begin with the simple case of only a single binary label per patient. The task is to learn a mapping from the vector X to y. This mapping is called the model and is performed by a learning algorithm such as stochastic gradient descent.
As discussed earlier, the degree to which the resulting model is causal is the degree to which it is an accurate representation of the true underlying process, denoted by f(x) in Figure 5-1. Depending on the data available, the degree of prior knowledge used in constraining the model’s structure, and the specific learning algorithm employed, we learn models that support differing degrees of causal interpretation.
Once the model is learned, given a new patient represented by a feature vector, we can then estimate the probability of the outcome. The data used to learn the model are called training data, and the new data used to assess how well a model
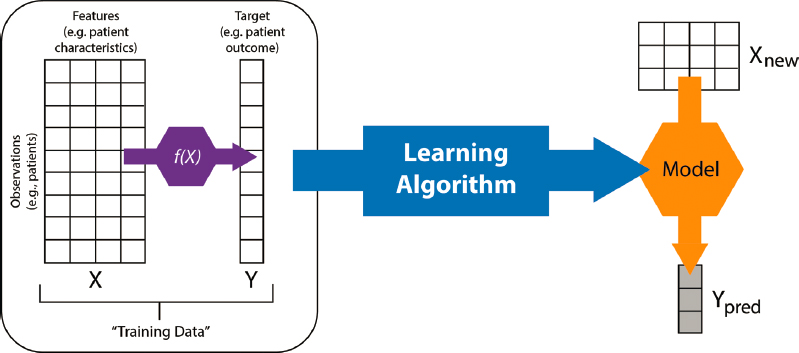
NOTE: A model is a map from inputs (X) to an output (y)—mathematically, a function. We implicitly assume that there is a real data-generating function, f(x), which is unknown and is what we are trying to represent at varying degrees of fidelity.
SOURCE: Developed by Alejandro Schuler and Nigam Shah for BIOMEDIN 215 at Stanford University.
performs are the test data (Wikipedia, 2019). Training data are often further split into training and validation subsets. Model selection, which is the selection of one specific model from among the many that are possible given the training data, is performed using the validation data.
Choosing the Data to Learn From
Bad data will result in bad models, recalling the age-old adage “garbage in, garbage out” (Kilkenny and Robinson, 2018). There is a tendency to hype AI as something magical that can learn no matter what the inputs are. In practice, the choice of data always trumps the choice of the specific mathematical formulation of the model.
In choosing the data for any model learning exercise, the outcome of interest (e.g., inpatient mortality) and the process for extracting it (e.g., identified using chart review of the discharge summary note) should be described in a reproducible manner. If the problem involves time-series data, the time at which an outcome is observed and recorded versus the time at which it needs to be predicted have to be defined upfront (see Figure 5-2). The window of data used to learn the model (i.e., observation window) and the amount of lead time needed from the prediction should be included.
It is necessary to provide a detailed description of the process of data acquisition, the criteria for subselecting the training data, and the description and prevalence of attributes that are likely to affect how the model will perform on a new dataset. For example, when building a predictive model, subjects in the training data may

NOTES: Specific events in the timeline are denoted by gray circles. The colored portions of the timeline below the gray line show the different types of data that may be collected at different encounters and the fact that not everything is collected at the same time. Almost no data source provides a continuous measurement of the patient’s health except data streams of intensive care unit monitors used in short stretches. (Wearables increasingly promise such continuous data but their use in health care is just beginning.) The red arrow shows a chosen point in the timeline where a prediction attempt is made. Only data prior to that are available for model learning for that prediction. Each prediction offers the chance of taking some action before the predicted event happens. The time interval between the prediction data and the soonest possible occurrence of the predicted event indicates the lead time available to complete the necessary mitigating action.
SOURCE: Developed by Alejandro Schuler and Nigam Shah for BIOMEDIN 215 at Stanford University.
not be representative of the target population (i.e., selection bias). Meanwhile, errors in measuring exposure or disease occurrences can be an important source of bias (i.e., measurement bias), especially when using EHR data as a source of measurements (Gianfrancesco et al., 2018). Both selection bias and measurement bias can affect the accuracy as well as generalizability of a predictive model learned from the data (Suresh and Guttag, 2019).
The degree to which the chosen data affect generalization of a model learned from it depends on the method used for modeling and the biases inherent in the data during their acquisition. For example, models can be susceptible to provider practice patterns; most models trained using supervised learning assume that practice patterns in the new environment are similar to those in the development environment (Schulam and Saria, 2017). The degree of left censoring, right censoring, or missingness can also affect generalization (Dyagilev and Saria, 2016; Molenberghs and Kenward, 2007; Schulam and Saria, 2018). Finally, the processes by which the data are generated and collected also change over time. This change, known as nonstationarity in the data, can have a significant effect on model performance (Jung and Shah, 2015). Using stale data can lead to suboptimal learning by models, which then get labeled as biased or unfair.
The decisions made during the creation and acquisition of datasets will be reflected in downstream models. In addition to knowing the final features representing patient data (see Figure 5-1), any preprocessing steps should be clearly documented and made available with the model. Such data-wrangling steps (e.g., how one dealt with missing values or irregularly sampled data) are often overlooked or not reported. The choices made around data preparation and transformation into the analytical data representation can contribute significantly to bias that then gets incorporated into the AI algorithm (Suresh and Guttag, 2019).
Often, the users of the model’s output hold the model itself responsible for such biases, rather than the underlying data and the model developer’s design decisions surrounding the data (Char et al., 2018). In nonmedical fields, there are numerous examples in which model use has reflected biases inherent in the data used to train them (Angwin et al., 2016; Char et al., 2018; O’Neil, 2017). For example, programs designed to aid judges in sentencing by predicting an offender’s risk of recidivism have shown racial discrimination (Angwin et al., 2016). In health care, attempts to use data from the Framingham Heart Study to predict the risk of cardiovascular events in minority populations have led to biased risk estimates (Gijsberts et al., 2015). Subtle discrimination inherent in health care delivery may be harder to anticipate; as a result, it may be more difficult to prevent an algorithm from learning and incorporating this type of bias (Shah et al., 2018). Such biases may lead to self-fulfilling prophesies: If clinicians always withdraw care from patients with certain findings (e.g., extreme prematurity or a brain injury), machine learning systems may conclude that such findings are always fatal. (Note that the degree to which such biases may affect actual patient care depends on the degree of causality ascribed to the model and to the process of choosing the downstream action.)
Learning Setup
In the machine learning literature, the dataset from which a model is learned is also called the training dataset. Sometimes a portion of this dataset may be set aside for tuning hyperparameters—the weights assigned to different variables and their combinations. This portion of the training data is referred to as the hyperparameter-validation dataset, or often just the validation dataset. The validation dataset confirms whether the choices of the values of the parameters in the model are correct or not. Note that the nomenclature is unfortunate, because these validation data have nothing to do with the notion of clinical validation or external validity.
Given that the model was developed from the training/validation data, it is necessary to evaluate its performance in classifying or making predictions on a “holdout” test set (see Figure 5-1). This test set is held out in the sense that it was not used to select model parameters or hyperparameters. The test set should be as
close as possible to the data that the model would be applied to in routine use. The choice of metrics used to assess a model’s performance is guided by the end goal of the modeling as well as the type of learning being conducted (e.g., unsupervised versus supervised). Here, we focus on metrics for supervised binary classifiers (e.g., patient risk stratification tools). The estimation of metrics can be obtained through cross validation where the whole dataset is split randomly into multiple parts with one part set as the test set and the remaining parts used for training a model.
Choosing Metrics of Model Performance
Recall and precision are two of the most debated performance metrics because they exhibit varying importance based on the use case. Sensitivity quantifies a classifier’s ability to identify the true positive cases. Typically, a highly sensitive classifier can reliably rule out a disease when its result is negative (Davidson, 2002). Precision quantifies a classifier’s ability to correctly identify a true positive case—that is, it estimates the number of times the classifier falsely categorizes a noncase as a case. Specificity quantifies the portion of actual negatives that are correctly identified as such. There is a trade-off between the recall, precision, and specificity measures, which needs to be resolved based on the clinical question of interest. For situations where we cannot afford to miss a case, high sensitivity is desired. Often, a highly sensitive classifier is followed up with a highly specific test to identify the false positives among those flagged by the sensitive classifier. The trade-off between specificity and sensitivity can be visually explored in the receiver operating characteristic (ROC) curve. The area under the ROC (AUROC) curve is the most popular index for summarizing the information in the ROC curves. When reporting results on the holdout test set, we recommend going beyond the AUROC curve and instead reporting the entire ROC curve as well as the sensitivity, specificity, positive predictive value, and negative predictive value at a variety of points on the curve that represent reasonable decision-making cutoffs (Bradley, 1997; Hanley and McNeil, 1982).
However, the limitations of the ROC curves are well known even though they continue to be widely used (Cook, 2007). Despite the popularity of AUROC curve and ROC curve for evaluating classifier performance, there are other important considerations. First, the utility offered by two ROC curves can be wildly different, and it is possible that classifiers with a lower overall AUROC curve have higher utility based on the shape of the ROC curve. Second, in highly imbalanced datasets, where negative and positive labels are not distributed equally, a precision-recall (PR) curve provides a better basis for comparing classifiers (McClish, 1989). Therefore, in order to enable meaningful comparisons, researchers should report both the AUROC curve and area under the PR curve, along with the actual curves, and error bars around the average classifier performance.
For decision making in the clinic, additional metrics such as calibration, net reclassification, and a utility assessment are necessary (Lorent et al., 2019; Shah et al., 2019; Steyerberg et al., 2010). While the ROC curves provide information about a classifier’s ability to discriminate a true case from a noncase, calibration metrics quantify how well the predicted probabilities of a true case being a case agree with observed proportions of cases and noncases. For a well-calibrated classifier, 90 of 100 samples with a predicted probability of 0.9 will be correctly identified true cases (Cook, 2008).
When evaluating the use of machine learning models, it is also important to develop parallel baselines, such as a penalized regression model applied on the same data that are supplied to more sophisticated models such as deep learning or random forests. Given the non-obvious relationship between a model’s positive predictive value, recall, and specificity to its utility, having these parallel models provides another axis of evaluation in terms of cost of implementation, interpretability, and relative performance.
Aside from issues related to quantifying the incremental value of using a model to improve care delivery, there are methodological issues in continuously evaluating or testing a model as the underlying data change. For example, a model for predicting 24-hour mortality could be retrained every week or every day as new data become available. It is unclear which metrics of the underlying data as well as of the model performance we should monitor to manage such continuously evolving models. It is also unclear how to set the retraining schedule, and what information should guide that decision. The issues of model surveillance and implementation are more deeply addressed in Chapter 6. Finally, there are unique regulatory issues that arise if a model might get retrained after it is approved and then behave differently, and some of the current guidance for these issues is discussed in Chapter 7.
DATA QUALIT Y
A variety of issues affect data integrity in health care. For example, the software for data retrieval, preprocessing, and cleaning is often lost or not maintained, making it impossible to re-create the same dataset. In addition, the data from the source system(s) may have been discarded or may have changed. The problem is further compounded by fast-changing data sources or changes over time in institutional data stores or governance procedures. Finally, silos of expertise and access around data sources create dependence on individual people or teams. When the collection and provenance of the data that a model is trained on is a black box, researchers must compensate with reliance on trusted individuals or teams, which is suboptimal and not sustainable in the long run. Developing AI based
on bad data further amplifies the potential negative impacts of poor-quality data. Consider, for example, that race and ethnicity information is simply not recorded, is missing, or is wrong in more than 30 to 40 percent of the records at most medical centers (Huser et al., 2016, 2018; Khare et al., 2017). Given the poor quality of these data, arguments about unfairness of predictive models for ethnic groups remain an academic discussion (Kroll, 2018). As a community, we need to address the quality of data that the vast majority of the enterprise is collecting. The quality is highly variable and acknowledging this variability as well as managing it during model building is essential. To effectively use AI, it is essential to follow good data practices in both the creation and curation of retrospective datasets for model training and in the prospective collection of the data. The quality of these data practices affects the development of models and the successful implementation at the point of care.
It is widely accepted that the successful development of an AI system requires high-quality data. However, the assessment of the quality of data that are available and the methodology to create a high-quality dataset are not standardized or often are nonexistent. Methods to assess data validity and reproducibility are often ad hoc. Efforts made by large research networks such as the Observational Health Data Science and Informatics collaborative as well as the Sentinel project have begun to outline quality assurance practices for data used to train AI models. Ideally, data should be cross-validated from multiple sources to best determine trustworthiness. Also, multiple subject matter experts should be involved in data validation (for both outcome and explanatory variables). In manually abstracted and annotated datasets, having multiple trained annotators can provide an accurate assessment of the ambiguity and variability inherent in data. For example, when tasked with identifying surgical site infection, there was little ambiguity whether infection was present or not; however, there was little agreement about the severity of the infection (Nuttall et al., 2016). Insufficiently capturing the provenance and semantics of such outcomes in datasets is at best inefficient. At worst, it can be outright dangerous, because datasets may have unspecified biases or assumptions, leading to models that produce inappropriate results in certain contexts. Ultimately, for the predictions (or classifications) from models to be trusted for clinical use, the semantics and provenance of the data used to derive them must be fully transparent, unambiguously communicated, and available for validation.
An often-missed issue around data is that the data used for training the model must be such that they are actually available in the real-world environment where the AI trained on the data will be used. For example, an AI analyzing electrocardiogram (ECG) waveforms must have a way to access the waveforms at the point of care. For instance, waveforms captured on a Holter monitor may not be available for clinical interpretation for hours, if not days, due to the difficulty of
processing the large amount of data, whereas an irregular heart rhythm presenting on a 12-lead ECG may be interpreted and acted upon within minutes. Therefore, AI development teams should have information technology (IT) engineers who are knowledgeable about the details of when and where certain data become available and whether the mechanics of data availability and access are compatible with the model being constructed.
Another critical point is that the acquisition of the data elements present in the training data must be possible without major effort. Models derived using datasets where data elements are manually abstracted (e.g., Surgical Risk Calculator from the American College of Surgeons) cannot be deployed without significant investment by the deploying site to acquire the necessary data elements for the patient for whom the model needs to be used. While this issue can be overcome with computational phenotyping methods, such methods struggle with portability due to EHR system variations resulting in different reporting schemes, as well as clinical practice and workflow differences. With the rise of interoperability standards such as the Fast Healthcare Interoperability Resource, the magnitude of this problem is likely to decrease in the near future. When computationally defined phenotypes serve as the basis for downstream analytics, it is important that computational phenotypes themselves be well managed and clearly defined and adequately reflect the target domain.
As a reasonable starting point for minimizing the data quality issues, data should adhere to the FAIR (findability, accessibility, interoperability, and reusability) principles in order to maximize the value of the data (Wilkinson et al., 2016). Researchers in molecular biology and bioinformatics put forth these principles, and, admittedly, their applicability in health care is not easy or straightforward.
One of the unique challenges (and opportunities) facing impactful design and implementation of AI in health care is the disparate data types that comprise today’s health care data. Today’s EHRs and wearable devices have greatly increased the volume, variety, and velocity of clinical data. The soon-to-be in-clinic promise of genomic data further complicates the problems of maintaining data provenance, timely availability of data, and knowing what data will be available for which patient at what time.
Always keeping a timeline view of the patient’s medical record is essential (see Figure 5-2), as is explicitly knowing the times at which the different data types across different sources come into existence. It stands to reason that any predictive or classification model operating at a given point in the patient timeline can only expect to use data that have come into being prior to the time at which the model is used (Jung et al., 2016; Panesar, 2019). Such a real-life view of data availability is crucial when building models, because using clean data gives an overly optimistic view of models’ performance and an unrealistic
impression of their potential value. Finally, we note that the use of synthetic data, if created to mirror real-life data in its missingness and acquisition delay by data type, can serve as a useful strategy for a model builder to create realistic training and testing environments for novel methods (Carnegie Mellon University, 2018; Franklin et al., 2017; Schuler, 2018).
EDUCATION
It is critical that we educate the community regarding data science, AI, medicine, and health care. Progress is contingent on creating a critical mass of experts in data science and AI who understand the mission, culture, workflow, strategic plan, and infrastructure of health care institutions.
As decision makers in health care institutions invest in data, tools, and personnel related to data science and AI, there is enormous pressure for rapid results. Such pressures raise two extremes of issues. On the one hand, the relative ease of implementing newly developed AI solutions rapidly can lead to the implementation of solutions in routine clinical care without an adequate understanding of their validity and potential influence on care, raising the potential for wasted resources and even patient harm (Herper, 2017). On the other hand, holding the AI models to superhuman standards and constantly requiring that evaluations outcompete doctors is also a flawed attitude that could lead to valuable solutions never getting implemented. Vendors of health care IT have an incentive to overstate the value of data science and AI generally. Limited attention has been given to the significant risk of harm, from wasting resources as well as from relying on evaluation strategies decoupled from the action they influence (Abrams, 2019) or relying on evaluation regimes that avoid simple and obvious baseline comparisons (Christodoulou et al., 2019).
KEY CONSIDERATIONS
The rapid increase in the volume and variety of data in health care has driven the current interest in the use of AI (Roski et al., 2014). There is active discussion and interest in addressing the potential ethical issues in using AI (Char et al., 2018), the need for humanizing AI (Israni and Verghese, 2019), the potential unintended consequences (Cabitza et al., 2017), and the need to tamper the hype (Beam and Kohane, 2018). However, more discovery and work in these areas is essential. The way that AI is developed, evaluated, and utilized in health care must change. At present, most of the existing discussion focuses on evaluating the model from a technical standpoint. A critically underassessed area is the net benefit of the integration of AI into clinical practice workflow (see Chapter 6).
Establishing Utility
When considering the use of AI in health care, it is necessary to know how one would act given a model’s output. While model evaluation typically focuses on metrics such as positive predictive value, sensitivity (or recall), specificity, and calibration, constraints on the action triggered by the model’s output (e.g., continuous rhythm monitoring constraint based on availability of Holter monitors) often can have a much larger influence in determining model utility (Moons et al., 2012). Completing model selection, then doing a net-benefit analysis, and later factoring work constraints are suboptimal (Shah et al., 2019). Realizing the benefit of using AI requires defining potential utility upfront. Only by including the characteristics of actions taken on the basis of the model’s predictions, and factoring in their implications, can a model’s potential usefulness in improving care be properly assessed.
Model Learning
After the potential utility has been established, model developers and model users need to interact closely during model learning because many modeling choices are dependent on the context of use of the model (Wiens et al., 2019). For example, the need for external validity depends on what one wishes to do with the model, the degree of agency ascribed to the model, and the nature of the action triggered by the model.
It is well known that biased data will result in biased models; thus, the data that are selected to learn from matter far more than the choice of the specific mathematical formulation of the model. Model builders need to pay closer attention to the data they train on and need to think beyond the technical evaluation of models. Even in technical evaluation, it is necessary to look beyond the ROC curves and examine multiple dimensions of performance (see Box 5-2). For decision making in the clinic, additional metrics such as calibration, net reclassification, and a utility assessment are necessary. Given the non-obvious relationship between a model’s positive predictive value, recall, and specificity to its utility, it is important to examine simple and obvious parallel baselines, such as a penalized regression model applied on the same data that are supplied to more sophisticated models such as deep learning.
The topic of interpretability deserves special discussion because of ongoing debates around interpretability, or the lack of it (Licitra et al., 2017; Lipton, 2016; Voosen, 2017). To the model builder, interpretability often means the ability to explain which variables and their combinations, in what manner, led to the output produced by the model (Friedler et al., 2019). To the clinical user, interpretability could mean one of two things: a sufficient enough understanding of what is going on, so that they can trust the output and/or be able to get liability insurance for its
recommendations; or enough causality in the model structure to provide hints as to what mitigating action to take. To avoid wasted effort, it is important to understand what kind of interpretability is needed in a particular application. A black box model may suffice if the output was trusted, and trust can be obtained by prospective assessment of how often the model’s predictions are correct and calibrated.
Data Quality
Bad data quality adversely affects patient care and outcomes (Jamal et al., 2009). A recent systematic review shows that the AI models could dramatically improve if four particular adjustments were made: use of multicenter datasets, incorporation of time-varying data, assessment of missing data as well as informative censoring, and development of metrics of clinical utility (Goldstein et al., 2017). As a reasonable starting point for minimizing the data quality issues, data should adhere to the FAIR principles in order to maximize the value of the data (Wilkinson et al., 2016). An often overlooked detail is when and where certain data become available and whether the mechanics of data availability and access are compatible with the model being constructed. In parallel, we need to educate the different stakeholders, and the model builders need to understand the datasets they learn from.
Stakeholder Education and Managing Expectations
The use of AI solutions presents a wide range of legal and ethical challenges, which are still being worked out (see Chapter 7). For example, when a physician makes decisions assisted by AI, it is not always clear where to place blame in the case of failure. This subtlety is not new to recent technological advancements, and in fact was brought up decades ago (Berg, 2010). However, most of the legal and ethical issues were never fully addressed in the history of computer-assisted decision support, and a new wave of more powerful AI-driven methods only adds to the complexity of ethical questions (e.g., the frequently condemned black box model) (Char et al., 2018).
The model builders need to better understand the datasets they choose to learn from. The decision makers need to look beyond technical evaluations and ask for utility assessments. The media needs to do a better job in articulating both immense potential and the risks of adopting the use of AI in health care. Therefore, it is important to promote a measured approach to adopting AI technology, which would further AI’s role as augmenting rather than replacing human actors. This framework could allow the AI community to make progress while managing evaluation challenges (e.g., when and how to employ interpretable models versus black box models) as well as ethical challenges that are bound to arise as the technology is widely adopted.
REFERENCES
Abdul, A., J. Vermeulen, D. Wang, B. Y. Lim, and M. Kankanhalli. 2018. Trends and trajectories for explainable, accountable and intelligible systems: An HCI research agenda. In Proceedings of the 2018 CHI Conference on Human Factors in Computing Systems. New York: ACM. https://dl.acm.org/citation.cfm?id=3174156 (accessed November 13, 2019).
Abrams, C. 2019. Google’s effort to prevent blindness shows AI challenges. The Wall Street Journal. https://www.wsj.com/articles/googles-effort-to-prevent-blindness-hits-roadblock-11548504004 (accessed November 13, 2019).
Amarasingham, R., R. E. Patzer, M. Huesch, N. Q. Nguyen, and B. Xie. 2014. Implementing electronic health care predictive analytics: Considerations and challenges. Health Affairs 33(7):1148–1154.
Angwin, J., J. Larson, S. Mattu, and L. Kirchner. 2016. Machine bias: There’s software used across the country to predict future criminals. And it’s biased against blacks. ProPublica. https://www.propublica.org/article/machine-bias-risk-assessments-in-criminal-sentencing (accessed November 13, 2019).
Beam, A. L., and I. S. Kohane. 2018. Big data and machine learning in health care. JAMA 319(13):1317–1318.
Berg, J. 2010. Review of “The Ethics of Consent, eds. Franklin G. Miller and Alan Wertheimer.” American Journal of Bioethics 10(7):71–72.
Bottou, L., F. E. Curtis, and J. Nocedal. 2018. Optimization methods for large-scale machine learning. SIAM Review 60(2):223–311.
Bradley, A. P. 1997. The use of the area under the ROC curve in the evaluation of machine learning algorithms. Pattern Recognition 30(7):1145–1159.
Breiman, L. 2001. Statistical modeling: The two cultures (with comments and a rejoinder by the author). Statistical Science 16(3):199–231.
Cabitza, F., R. Rasoini, and G. F. Gensini. 2017. Unintended consequences of machine learning in medicine. JAMA 318(6):517–518.
Carnegie Mellon University. 2018. Atlantic Causal Inference Conference—2018: Data Challenge. https://www.cmu.edu/acic2018/data-challenge/index.html (accessed November 13, 2019).
Char, D. S., N. H. Shah, and D. Magnus. 2018. Implementing machine learning in health care—Addressing ethical challenges. New England Journal of Medicine 378(11):981–983.
Chen, I. Y., P. Szolovits, and M. Ghassemi. 2019. Can AI help reduce disparities in general medical and mental health care? AMA Journal of Ethics 21(2):167–179.
Christodoulou, E., J. Ma, G. S. Collins, E. W. Steyerberg, J. Y. Verbakel, and B. Van Calster. 2019. A systematic review shows no performance benefit of machine
learning over logistic regression for clinical prediction models. Journal of Clinical Epidemiology 110:12–22.
Cleophas, T. J., A. H. Zwinderman, and H. I. Cleophas-Allers. 2013. Machine learning in medicine. New York: Springer. https://www.springer.com/gp/book/9789400758230 (accessed November 13, 2019).
Cook, N. R. 2007. Use and misuse of the receiver operating characteristic curve in risk prediction. Circulation 115(7):928–935.
Cook, N. R. 2008. Statistical evaluation of prognostic versus diagnostic models: Beyond the ROC curve. Clinical Chemistry 54(1):17–23.
Cushman, W. C., P. K. Whelton, L. J. Fine, J. T. Wright, Jr., D. M. Reboussin, K. C. Johnson, and S. Oparil. 2016. SPRINT trial results: Latest news in hypertension management. Hypertension 67(2):263–265.
Davidson, M. 2002. The interpretation of diagnostic tests: A primer for physiotherapists. Australian Journal of Physiotherapy 48(3):227–232.
Doshi-Velez, F., Y. Ge, and I. Kohane. 2014. Comorbidity clusters in autism spectrum disorders: An electronic health record time-series analysis. Pediatrics 133(1):54–63.
Dyagilev, K., and S. Saria. 2016. Learning (predictive) risk scores in the presence of censoring due to interventions. Machine Learning 102(3):323–348.
Esteva, A., A. Robicquet, B. Ramsundar, V. Kuleshov, M. DePristo, K. Chou, C. Cui, G. Corrado, S. Thrun, and J. Dean. 2019. A guide to deep learning in healthcare. Nature Medicine 25(1):24.
Fox, C. S. 2010. Cardiovascular disease risk factors, type 2 diabetes mellitus, and the Framingham Heart Study. Trends in Cardiovascular Medicine 20(3):90–95.
Franklin, J. M., W. Eddings, P. C. Austin, E. A. Stuart, and S. Schneeweiss. 2017. Comparing the performance of propensity score methods in healthcare database studies with rare outcomes. Statistics in Medicine 36(12):1946–1963.
Friedler, S. A., C. D. Roy, C. Scheidegger, and D. Slack. 2019. Assessing the local interpretability of machine learning models. arXiv.org. https://ui.adsabs.harvard.edu/abs/2019arXiv190203501S/abstract (accessed November 13, 2019).
Gianfrancesco, M., S. Tamang, and J. Yazdanzy. 2018. Potential biases in machine learning algorithms using electronic health record data. JAMA Internal Medicine 178(11):1544–1547.
Gijsberts, C. M., K. A. Groenewegen, I. E. Hoefer, M. J. Eijkemans, F. W. Asselbergs, T. J. Anderson, A. R. Britton, J. M. Dekker, G. Engström, G. W. Evans, and J. De Graaf. 2015. Race/ethnic differences in the associations of the Framingham risk factors with carotid IMT and cardiovascular events. PLoS One 10(7):e0132321.
Goldstein, B. A., A. M. Navar, M. J. Pencina, and J. Ioannidis. 2017. Opportunities and challenges in developing risk prediction models with electronic health
records data: A systematic review. Journal of the American Medical Informatics Association 24(1):198–208.
Gottesman, O., F. Johansson, M. Komorowski, A. Faisal, D. Sontag, F. Doshi-Velez, and L. A. Celi. 2019. Guidelines for reinforcement learning in healthcare. Nature Medicine 25(1): 16–18.
Goyal, D., D. Tjandra, R. Q. Migrino, B. Giordani, Z. Syed, J. Wiens, and Alzheimer’s Disease Neuroimaging Initiative. 2018. Characterizing heterogeneity in the progression of Alzheimer’s disease using longitudinal clinical and neuroimaging biomarkers. Alzheimer’s & Dementia: Diagnosis, Assessment & Disease Monitoring 10:629–637.
Greenland, P., S. C. Smith, Jr., and S. M. Grundy. 2001. Improving coronary heart disease risk assessment in asymptomatic people: Role of traditional risk factors and noninvasive cardiovascular tests. Circulation 104(15):1863–1867.
Hanley, J. A., and B. J. McNeil. 1982. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143(1):29–36.
He, J., S. L. Baxter, J. Xu, J. Xu, X. Zhou, and K. Zhang. 2019. The practical implementation of artificial intelligence technologies in medicine. Nature Medicine 25(1):30–36.
Herasevich, V., B. Pickering, and O. Gajic. 2018. How Mayo Clinic is combating information overload in critical care units. Harvard Business Review. https://www.bizjournals.com/albany/news/2018/04/23/hbr-how-mayo-clinic-is-combating-information.html (accessed November 13, 2019).
Herper, M. 2017. MD Anderson benches IBM Watson in setback for artificial intelligence in medicine. Forbes. https://www.forbes.com/sites/matthewherper/2017/02/19/md-anderson-benches-ibm-watson-in-setback-for-artificial-intelligence-in-medicine/#7a051f7f3774 (accessed November 13, 2019).
Huser, V., F. J. DeFalco, M. Schuemie, P. B. Ryan, N. Shang, M. Velez, R. W. Park, R. D. Boyce, J. Duke, R. Khare, and L. Utidjian. 2016. Multisite evaluation of a data quality tool for patient-level clinical data sets. EGEMS (Washington, DC) 4(1):1239.
Huser, V., M. G. Kahn, J. S. Brown, and R. Gouripeddi. 2018. Methods for examining data quality in healthcare integrated data repositories. Pacific Symposium on Biocomputing. 23:628–633.
Hutson, M. 2017. This computer program can beat humans at Go—with no human instruction. Science Magazine. https://www.sciencemag.org/news/2017/10/computer-program-can-beat-humans-go-no-human-instruction (accessed November 13, 2019).
Israni, S. T., and A. Verghese. 2019. Humanizing artificial intelligence. JAMA 321(1):29–30.
Jamal, A., K. McKenzie, and M. Clark. 2009. The impact of health information technology on the quality of medical and health care: A systematic review. Health Information Management Journal 38(3):26–37.
Jung, K., and N. H. Shah. 2015. Implications of non-stationarity on predictive modeling using EHRs. Journal of Biomedical Informatics 58:168–174.
Jung, K., S. Covington, C. K. Sen, M. Januszyk, R. S. Kirsner, G. C. Gurtner, and N. H. Shah. 2016. Rapid identification of slow healing wounds. Wound Repair and Regeneration 24(1):181–188.
Ke, C., Y. Jin, H. Evans, B. Lober, X. Qian, J. Liu, and S. Huang. 2017. Prognostics of surgical site infections using dynamic health data. Journal of Biomedical Informatics 65:22–33.
Khare, R., L. Utidjian, B. J. Ruth, M. G. Kahn, E. Burrows, K. Marsolo, N. Patibandla, H. Razzaghi, R. Colvin, D. Ranade, and M. Kitzmiller. 2017. A longitudinal analysis of data quality in a large pediatric data research network. Journal of the American Medical Informatics Association 24(6):1072–1079.
Kilkenny, M. F., and K. M. Robinson. 2018. Data quality: “Garbage in–garbage out.” Health Information Management 47(3):103–105.
Komorowski, M., L. A. Celi, O. Badawi, A. C. Gordon, and A. A. Faisal. 2018. The Artificial Intelligence Clinician learns optimal treatment strategies for sepsis in intensive care. Nature Medicine 24(11):1716–1720.
Kroll, J. A. 2018. Data science data governance [AI Ethics]. IEEE Security & Privacy 16(6):61–70.
Kwong, C., A. Y. Ling, M. H. Crawford, S. X. Zhao, and N. H. Shah. 2017. A clinical score for predicting atrial fibrillation in patients with crytogenic stroke or transient ischemic attack. Cardiology 138(3):133–140.
Licitra, L., A. Trama, and H. Hosni. 2017. Benefits and risks of machine learning decision support systems. JAMA 318(23):2356.
Lipton, Z. C. 2016. The mythos of model interpretability. arXiv.org. https://arxiv.org/abs/1606.03490 (accessed November 13, 2019).
Lorent, M., H. Maalmi, P. Tessier, S. Supiot, E. Dantan, and Y. Foucher. 2019. Meta-analysis of predictive models to assess the clinical validity and utility for patient-centered medical decision making: Application to the Cancer of the Prostate Risk Assessment (CAPRA). BMC Medical Informatics and Decision Making 19(1):Art. 2.
Mason, P. K., D. E. Lake, J. P. DiMarco, J. D. Ferguson, J. M. Mangrum, K. Bilchick, L. P. Moorman, and J. R. Moorman. 2012. Impact of the CHA2DS2-VASc score on anticoagulation recommendations for atrial fibrillation. American Journal of Medicine 125(6):603.e1–603.e6.
McClish, D. K. 1989. Analyzing a portion of the ROC curve. Medical Decision Making 9(3):190–195.
Meskó, B., G. Hetényi, and Z. Győrffy. 2018. Will artificial intelligence solve the human resource crisis in healthcare? BMC Health Services Research 18(1):Art. 545.
Molenberghs, G., and M. Kenward. 2007. Missing data in clinical studies, vol. 61. Hoboken, NJ: John Wiley & Sons. https://www.wiley.com/en-us/Missing+Data+in+Clinical+Studies-p-9780470849811 (accessed November 13, 2019).
Moons, K. G., A. P. Kengne, D. E. Grobbee, P. Royston, Y. Vergouwe, D. G. Altman, and M. Woodward. 2012. Risk prediction models: II. External validation, model updating, and impact assessment. Heart 98(9):691–698.
Nuttall, J., N. Evaniew, P. Thornley, A. Griffin, B. Deheshi, T. O’Shea, J. Wunder, P. Ferguson, R. L. Randall, R. Turcotte, and P. Schneider. 2016. The inter-rater reliability of the diagnosis of surgical site infection in the context of a clinical trial. Bone & Joint Research 5(8):347–352.
O’Neil, C. 2017. Weapons of math destruction: How big data increases inequality and threatens democracy. New York: Broadway Books.
Panesar, A., 2019. Machine learning and AI for healthcare: Big data for improved health outcomes. New York: Apress.
Pearl, J., and D. Mackenzie. 2018. The book of why: The new science of cause and effect. New York: Basic Books.
Poursabzi-Sangdeh, F., D. G. Goldstein, J. M. Hofman, J. W. Vaughan, and H. Wallach. 2018. Manipulating and measuring model interpretability. arXiv.org. https://arxiv.org/abs/1802.07810 (accessed November 13, 2019).
Rajkomar, A., E. Oren, K. Chen, A. M. Dai, N. Hajaj, M. Hardt, P. J. Liu, X. Liu, J. Marcus, M. Sun, P. Sundberg, H. Yee, K. Zhang, Y. Zhang, G. Flores, G. E. Duggan, J. Irvine, Q. Le, K. Litsch, A. Mossin, J. Tansuwan, D. Wang, J. Wexler, J. Wilson, D. Ludwig, S. L. Volchenboum, K. Chou, M. Pearson, S. Madabushi, N. H. Shah, A. J. Butte, M. D. Howell, C. Cui, G. S. Corrado, and J. Dean. 2018. Scalable and accurate deep learning with electronic health records. NPJ Digital Medicine 1(1):18.
Rajpurkar, P., J. Irvin, R. L. Ball, K. Zhu, B. Yang, H. Mehta, T. Duan, D. Ding, A. Bagul, C. P. Langlotz, B. N. Patel, K. W. Yeom, K. Shpanskaya, F. G. Blankenberg, J. Seekins, T. J. Amrhein, D. A. Mong, S. S. Halabi, E. J. Zucker, A. Y. Ng, and M. P. Lungren. 2018. Deep learning for chest radiograph diagnosis: A retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Medicine 15(11):e1002686.
Roski, J., G. Bo-Linn, and T. Andrews. 2014. Creating value in healthcare through big data: Opportunities and policy implications. Health Affairs 33(7):1115–1122.
Sachs, K., O. Perez, D. Pe’er, D. A. Lauffenburger, and G. P. Nolan. 2005. Causal protein-signaling networks derived from multiparameter single-cell data. Science 308(5721):523–529.
Saria, S. 2018. Individualized sepsis treatment using reinforcement learning. Nature Medicine 24(11):1641–1642.
Saria, S., and A. Goldenberg. 2015. Subtyping: What it is and its role in precision medicine. IEEE Intelligent Systems 30(4):70–75.
Schulam, P., and S. Saria. 2017. Reliable decision support using counterfactual models. arXiv.org. https://arxiv.org/abs/1703.10651 (accessed November 13, 2019).
Schulam, P., and S. Saria. 2018. Discretizing logged interaction data biases learning for decision-making. arXiv.org. https://arxiv.org/abs/1810.03025 (accessed November 13, 2019).
Schuler, A. 2018. Some methods to compare the real-world performance of causal estimators. Ph.D. dissertation, Stanford University. http://purl.stanford.edu/vg743rx0211 (accessed November 13, 2019).
Shah, N. D., E. W. Steyerberg, and D. M. Kent. 2018. Big data and predictive analytics: Recalibrating expectations. JAMA 320(1):27–28.
Shah, N. H., A. Milstein, and S. C. Bagley. 2019. Making machine learning models clinically useful. JAMA 322(14):1351–1352. https://doi.org/10.1001/jama.2019.10306.
Shah, S. J., D. H. Katz, S. Selvaraj, M. A. Burke, C. W. Yancy, M. Gheorghiade, R. O. Bonow, C. C. Huang, and R. C. Deo. 2015. Phenomapping for novel classification of heart failure with preserved ejection fraction. Circulation 131(3):269–279.
Shortliffe, E. H. 1974. MYCIN: A rule-based computer program for advising physicians regarding antimicrobial therapy selection. No. AIM-251. Stanford University. https://searchworks.stanford.edu/view/12291681 (accessed November 13, 2019).
Sohn, S., D. W. Larson, E. B. Habermann, J. M. Naessens, J. Y. Alabbad, and H. Liu. 2017. Detection of clinically important colorectal surgical site infection using Bayesian network. Journal of Surgical Research 209:168–173.
Steyerberg, E. W., A. J. Vickers, N. R. Cook, T. Gerds, M. Gonen, N. Obuchowski, M. J. Pencina, and M. W. Kattan. 2010. Assessing the performance of prediction models: A framework for some traditional and novel measures. Epidemiology 21(1):128–138.
Subbaswamy, A., P. Schulam, and S. Saria. 2018. Learning predictive models that transport. arXiv.org. https://arxiv.org/abs/1812.04597 (accessed November 13, 2019).
Suresh, H., and J. V. Guttag. 2019. A framework for understanding unintended consequences of machine learning. arXiv.org. https://arxiv.org/abs/1901.10002 (accessed November 13, 2019).
Vellido, A. 2019. The importance of interpretability and visualization in machine learning for applications in medicine and health care. Neural Computing and Applications. https://doi.org/10.1007/s00521-019-04051-w (accessed November 13, 2019).
Verghese, A., N. H. Shah, and R. A. Harrington. 2018. What this computer needs is a physician: Humanism and artificial intelligence. JAMA 319(1):19–20.
Voosen, P. 2017. The AI detectives. Science 357(6346):22–27. https://science.sciencemag.org/content/357/6346/22.summary (accessed November 13, 2019).
Wang, W., M. Bjarnadottir, and G. G. Gao. 2018. How AI plays its tricks: Interpreting the superior performance of deep learning-based approach in predicting healthcare costs. SSRN. https://papers.ssrn.com/sol3/papers.cfm?abstract_id=3274094 (accessed November 13, 2019).
Wiens, J., S. Saria, M. Sendak, M. Ghassemi, V. X. Liu, F. Doshi-Velez, K. Jung, K. Heller, D. Kale, M. Saeed, P. N. Ossocio, S. Thadney-Israni, and A. Goldenberg. 2019. Do no harm: A roadmap for responsible machine learning for health care. Nature Medicine 25(9):1337–1340.
Wikipedia. 2019. Training, validation, and test sets. https://en.wikipedia.org/wiki/Training,_validation,_and_test_sets (accessed November 13, 2019).
Wilkinson, M. D., M. Dumontier, I. J. Aalbersberg, G. Appleton, M. Axton, A. Baak, N. Blomberg, J. W. Boiten, L. B. da Silva Santos, P. E. Bourne, and J. Bouwman. 2016. The FAIR Guiding Principles for scientific data management and stewardship. Scientific Data 3:Art 160018.
Williams, J. B., D. Ghosh, and R. C. Wetzel. 2018. Applying machine learning to pediatric critical care data. Pediatric Critical Care Medicine 19(7):599–608.
Wu, A. 2019. Solving healthcare’s big epidemic—Physician burnout. Forbes. https://www.forbes.com/sites/insights-intelai/2019/02/11/solving-healthcares-big-epidemicphysician-burnout/#64ed37e04483 (accessed November 13, 2019).
Yadlowsky, S., R. A. Hayward, J. B. Sussman, R. L. McClelland, Y. I. Min, and S. Basu. 2018. Clinical implications of revised pooled cohort equations for estimating atherosclerotic cardiovascular disease risk. Annals of Internal Medicine 169(1):20–29.
Yu, K. H., A. L. Beam, and I. S. Kohane. 2018. Artificial intelligence in healthcare. Nature Biomedical Engineering 2(10):719–731.
Zhan, A., S. Mohan, C. Tarolli, R. B. Schneider, J. L. Adams, S. Sharma, M. J. Elson, K. L. Spear, A. M. Glidden, M. A. Little, and A. Terzis. 2018. Using smartphones and machine learning to quantify Parkinson disease severity: The mobile Parkinson disease score. JAMA Neurology 75(7):876–880.
Zhou, Z. H. 2017. A brief introduction to weakly supervised learning. National Science Review 5(1):44–53.
Suggested citation for Chapter 5: Liu, H., H. Estiri, J. Wiens, A. Goldenberg, S. Saria, and N. Shah. 2020. Artificial intelligence model development and validation. In Artificial intelligence in health care: The hope, the hype, the promise, the peril. Washington, DC: National Academy of Medicine.



























