Microbial and Phenotypic Definition of Rats and Mice: Proceedings of the 1998 US/Japan Conference (1999)
Chapter: Genetic and Phenotypic Definition of Laboratory Mice and Rats / What Constitutes an Acceptable Genetic-Phenotypic Definition
Common Disease Models
Development of the modern mapping techniques described above has also made it possible to map multiple genes causing common adulthood model diseases in mice (for example, diabetes in non-obese diabetes [NOD] strain). From those studies, it is assumed that although most mutations have mild effects, a specific combination of them can facilitate the expression of an ethnological mutation. Because common adulthood (life-style) diseases such as diabetes and cancer appear to be caused by the specific combination of many normal variant genes and, in many cases, etiological genes, the animal models for them should replicate human disease states. A broader study of gene loci related to diseases requires more variant alleles in mice for analyzing the molecular mechanism of gene manifestation. Asian mice are useful for that purpose because they are genetically more remote from laboratory mice and have plenty of variant alleles. We were able to conduct a DNA analysis using 60 marker DNA loci with Asian mice (Moriwaki and others 1999).
The finding that variant genes contained in the Asian wild mice sometimes have a long evolutionary history is biologically important to investigate the mechanism of gene function. It is not possible to select for long evolutionary history in fancy mice and laboratory mice.
As seen in the NOD experiment conducted by Wakana and colleagues (Wakana and others 1997), genetic introduction of a genetically remote allele of Idd-4 in Asian wild-derived MSM strain (established from wild mice collected in Mishima) exhibited a significant increase in frequency of diabetes. This strain should be a useful model to analyze Idd-4 function, which cannot be observed by the introduction of BALB/c or C57BL/6 alleles.
Another example of the characteristic function of Asian wild-derived alleles is the expression of the Rim4 mutant phenotype, polydactyly, which was completely suppressed in the Asian wild-derived genetic background (Masuya and others 1997). One might expect some "dominant negative" structural change in the gene product.
Conclusion
Animal models of common adulthood diseases such as diabetes and cancer have indicated that these diseases are apparently caused by the specific combination of many normal variant genes and possibly some etiological genes. To further our knowledge requires additional animal models so that we can identify a large number of variant alleles that vary within the normal range. For this purpose, Asian wild-derived genes are useful not only for the number of variations, but also for the large differences in the genome structure, which sometimes give rise to a "dominant negative" effect. These characteristics are useful for analyzing the mechanism of normal gene functions as seen in the case of Rim4 mouse.
References
Bailey, D. W. 1971. Recombinant-inbred strains. Transplantation 11:325–327.
Festing, M. F., A. Yang, and A. M. Malkinson. 1994. At least four genes and sex are associated with susceptibility to urethane-induced pulmonary adenoma in mice. Genet. Res. 64:99–106.
Goodenough, U., and R. P. Levine. 1974. Genetics. Holt, Rinehart and Winston, Inc., New York.
Manenti, G., M. Galibordi, R. Elango, A. Fiorino, L. De-Gregorio, F. S. Falvella, K. Hunter, D. Housman, M. A. Pierotti, and T. A. Dragani. 1996. Genetic mapping of a pulmonary adenoma resistance (Par 1) in mouse. Nat. Genet. 12:455–457.
Masuya, H. T. Sagai, K. Moriwaki, and T. Shiroishi. 1997. Multigenic control of the localization of the zone of polarizing activity in limb morphogenesis in the mouse. Dev. Biol. 182:42–51.
Moriwaki, K., N. Miyashita, Y. Yamaguchi, and T. Shiroishi. 1999. Multiple genes governing biological functions in the genetic backgrounds of laboratory mice and Asian wild mice. Prog. Exp. Tumor Res. Karger, Basel. 30:1–12.
Nishimura, M., N. Hirayama, T. Serikawa, K. Kanehira, Y. Matsushima, H. Katoh, S. Wakana, A. Kojima, and H. Hiai. 1995. The SMZA: A new set of recombinant inbred strain of mice consisting of 26 substrains and their genetic profile. Mamm. Genome 6:850–857.
Obata, M., H. Nishimori, K. Ogawa, and G. H. Lee. 1996. Identification of the Par2 (pulmonary adenoma resistance) locus on mouse chromosome 18, a major genetic determinant for lung carcinogen resistance in BALB/cByJ mice. Oncogene 13:1599–1604.
Pataer, A., M. Nishimura, T. Kamoto, K. Ichioka, M. Sato, and H. Hiai. 1997. Genetic resistance to urethane-induced pulmonary adenomas in SMXA recombinant inbred mouse strains. Cancer Res. 57:2904–2908.
Sibilia, M., and E. F. Wagner. 1995. Strain-dependent epithelial defects in mice lacking the EGF receptor. Science 269:234–238.
Silver, L. M. 1995. Mouse Genetics. Oxford University Press, New York.
Snell, G., J. Dausset, and S. Nathenson. 1976. Histocompatibility. Academic Press, New York.
Threadgill, D. W., A. A. Dlugosz, L. A. Hansen, T. Tennenbaum, U. Lichti, D. Yee, C. LaMantia, T. Mourton, K. Herrup, R. C. Harris, J. A. Barnard, S. H. Yuspa, R. J. Coffey, and T. Magnuson. 1995. Targeted disruption of mouse EGF receptor: Effect of genetic background on mutant phenotype. Science 269:230–234.
Wakana, S., T. Shiroishi, K. Moriwaki, A. Kono, and T. Nomura. 1997. Susceptibility gene Idd-4 controls onset of IDDM: An allele from the nondiabetic MSM strain is associated with early onset of diabetes in mice. 11th Annual Mouse Genome Conference, Miami, Florida (Abstract).
Wolf, C. R., and C. J. Hendérson. 1998. Use of transgenic animals in understanding molecular mechanisms of toxicity. J. Pharm. Pharmacol. 50:567–574.
Genetic and Phenotypic Definition of Laboratory Mice and Rats / What Constitutes an Acceptable Genetic-Phenotypic Definition
Hideki Katoh
Chief, Genetics Division
ICLAS Monitoring Center
Central Institute for Experimental Animals
Kanagawa, Japan
Genetic Studies on Closed Colonies of the Rat
The following questions are important to all of us who are interested in human racial differences:
- Are the ancestors of present races common or different?
- How do we understand genetic relations among the races?
Population genetics studies provide answers to these questions as follows.
Human Ancestry
We know that all human races have one common ancestor, whose progeny migrated across the world roughly 50,000 to 200,000 years ago. After these common ancestors settled in various areas and adapted to the environments, they developed as races.
That today's races have a common ancestor means that they had common genes. These races exist as a result of accumulated genetic changes caused by gene mutations and of gene frequencies at many loci. Such genetic changes resulted in racial diversity, with different genetic characteristics such as skin color and height.
Genetic Relations Among Races
Population geneticists have calculated gene frequencies of blood types and have shown similarities among races using differences of various blood type frequencies from blood group data of all of the races. Genes and their frequencies of common ancestors of the human races were randomly changed and varied when the ancestors dispersed to various parts of the world and settled there. Randomly occurring changes of gene frequency are called genetic drift. One example of genetic drift obtained by computer simulation can be seen in Figure 1. If the ancestors had A and B blood types at rates of 50% each, one of the two races will show a higher percentage of the A type and the other loses the A type after 300 generations. These changes are natural and occur by chance. Assuming a breeding cycle of 30 years, 300 generations is equivalent to about 10,000 years in humans. This time is sufficient to develop races different from the original one. However, it is important to remember that mating between races is possible, and such offspring will also be reproductive. This ability to produce fertile offspring means that no race has developed into subspecies. In the mouse, 1,000,000 years are required for genetic divergence to lead to a subspecies.
Ethnic Differences
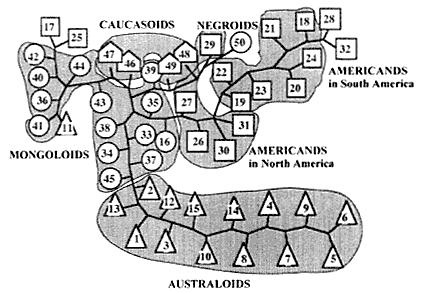
It is interesting to consider the number of loci required to identify ethnic differences of human races. In Figure 2, the results for several racial groups can be seen using 12 loci, with Caucasoids, Mongoloids, Negroids, and Australoids clearly identified. These results agree with those of anthropology and cultural anthropology. What happens if we have fewer loci for an ethnic analysis? Figure 3
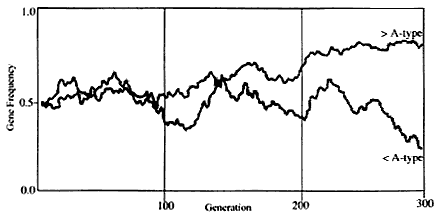
Figure 1
A computer-simulated result of genetic drift.
illustrates the results for the same racial groups using only three loci. As expected, the results are incomplete because some races are grouped into the wrong groups. Thus more reliable results are obtained in a population genetics study by using as many loci as possible.
Laboratory Animals
Closed Colonies
To apply the genetic information described above to laboratory animals, it is necessary to define ''laboratory animals.'' Mice and rats include both laboratory and wild animals. Wild animals have been domesticated, and inbred strains and closed colonies have been bred as laboratory animals. In addition, there are many inbred strains and closed colonies with particular characteristics. As with humans, laboratory and wild animals also have common ancestors and so have inherited the same set of genes a long time ago.
At the beginning of the 20th century, inbreeding was started to obtain inbred lines of mice. The oldest lines in the mouse have approximately an 80-year history. Because mice and rats reproduce three times (generations) a year, an 80-year history is equivalent to 240 generations. We can anticipate that two lines derived from a single ancestor differ genetically in the same way as two human races differ. It is important to remember that in spite of the two lines being separated 80 years ago, they can still mate and produce fertile offspring.
However, in the case of closed colonies, mice and rats have approximately a 70-year history (Lindsey 1979; Morse 1981). Closed colonies also have several sublines derived from the main colonies. Genetic analysis reveals fewer differences among closed colonies of mice and rats than among humans, indicating less genetic drift.
One important problem exists in the case of laboratory animals, as illustrated in the following extreme example. Suppose that a colony of breeder A was supplied to breeder B 10 years ago. Genetic drift is sufficiently slow that we can assume that in 10 years, the two lines will not diverge dramatically. In fact, the two breeders do anticipate such changes. However, it is possible that breeding schemes can be accidentally mixed up by people working in animal facilities. It is possible that they will take males and females that are closely related, thereby establishing a subcolony in a breeding facility. In such a case, the two colonies will be genetically quite different, and a typical bottleneck effect will occur.
Consider a more ideal situation in a rat colony. Suppose that a researcher is working on allotransplantation of a cancer cell line using a closed colony. Assume that the cancer has the A-antigen of RT1. In the closed colony as a recipient of the cancer, the percentages of the animals with A, AB, and B were 25%, 50%, and 25% in one generation. It can be assumed that the cancer will be transplantable in 75% of the animals with the A-antigen on average. If animals with the
B-type contributed to the subsequent generation of the divided colony, they will produce only offspring carrying B, which will reject the cancer. Such cases generally do not arise, but the worse case (such as artificially lowering the frequency of A-antigen-positive animals) should be carefully considered.
Demonstration of the Existence of Closed Colonies Using Genetic Monitoring
Closed colonies of rats are known to show genetic polymorphisms at many loci. When they are maintained under unplanned mating, the stocks may show genetic instability. One cause of instability is a bottleneck effect caused by caesarian section, followed by an increase in an inbreeding coefficient in the colony. In this case, the number of loci showing genetic polymorphisms decreases with each generation.
Generally, genetic variation in a closed colony is greater than that seen with an inbred strain. When a colony has been genetically altered for any of the reasons described above, it will be difficult for users of the stocks to interpret their data because genetic changes have occurred. Therefore, we should periodically test closed colonies using genetic methods to confirm genetic stability.
From the viewpoint of population genetics, we studied closed colonies of rats for 3 years (1988 to 1990). The Ministry of Education, Science, Culture, and Sports of Japan provided a grant for our study in which we performed genetic analysis on closed colonies of rats with the following aims: (1) to obtain genetic profiles of closed colonies of rats, (2) to obtain monitoring profiles of closed colonies, and (3) to determine specific genes identifying each colony. As mentioned above, an additional goal of this study was to establish methods of genetic monitoring that are appropriate for closed colonies.
Materials and Methods
As shown in Figure 4, six closed colonies were obtained from four breeders in Japan: Jcl:Wistar, Crj:Wistar, Iar:Wistar, Jcl:SD, Crj:SD, and Nr:Donryu. Sixty rats consisting of 30 females and 30 males of each colony were used for genetic study. Of the genetic markers used in the study, 21 are biochemical markers and one is an immunological marker (Table 1).
Results
Gene frequencies and specific markers identifying colony.
Gene frequencies of 21 biochemical markers are shown in Table 2. Results for RT1 are summarized in Table 3. Markers showing > 0.017 genotypic frequency (equivalent to at least one occurrence in 60 animals tested) were counted. Markers identifying colonies were demonstrated as follows: Acon1a: SD stocks; Ahd2b:
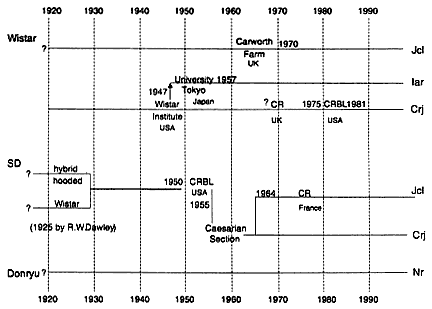
Figure 4
History of the five outbred stocks of the rat used in this study.
Jcl:Wistar; Amylb: Crj:SD; Es3b: Iar:Wistar; Es4c: Nr:Donryu; RT1.Ak: Iar:Wistar; RT1.Ed: Crj:Wistar.
Average heterozygosity.
As shown in Table 4, percentages of polymorphic loci in six colonies variéd from 9.5% (Jcl:Wistar) to 61.9% (SD), and average heterozygosities of six colonies ranged from 0.007 (Donryu) to 0.251 (Crj:SD). Because average heterozygosities of mice and human are 0.078 and 0.099, respectively (Nei 1987), those of rat closed colonies were reasonable. Jcl:Wistar revealed the lowest percentage (9.5%) of polymorphic loci. The breeder stated that this colony was reconstituted from several pairs by cesarian section to produce a SPF colony, which might have led to elevation of homozygosity at most loci. Donryu was second in percentage of polymorphic loci and first for average heterozygosity. This ranking was caused by a significant imbalance of allele frequencies of polymorphic loci in the Donryu colony.
Genetic distance.
In Table 5 are shown the genetic distances among the colonies. Each value represents the level of genetic difference between two colonies. The lowest value was 0.151 between Jcl:SD and Crj:SD, and the highest one was 0.474 between Jcl:SD and Donryu. The close relation between Iar:Wistar and Donryu is shown by a distance value of 0.160.
TABLE 1 Marker Loci, Samples and Testing Methods
|
Gene Symbol |
Gene Name |
Samples |
Testing Methods |
|
|
Biochemical markers |
||||
|
1 |
Acon1 |
Aconitase 1 |
Kidney |
CAE |
|
2 |
Ahd2 |
Aldehyde dehydrogenase 2 |
Liver |
IEF (pH3.5–10) |
|
3 |
Ahdc |
Aldehyde dehydrogenase c |
Liver |
IEF (pH3.5–10) |
|
4 |
Akp1 |
Alkaline phosphatase 1 |
Kidney |
IEF (pH5–7) |
|
5 |
Alp1 |
Serum alkaline phosphatase 1 |
Kidney |
IEF (pH5–7) |
|
6 |
Amy1 |
Amylase 1 |
Pancreas |
CAE |
|
7 |
Es1 |
Esterase 1 |
Plasma |
CAE |
|
8 |
Es2 |
Esterase 2 |
Plasma |
PAGE (10% gel) |
|
9 |
Es3 |
Esterase 3 |
Small intestine |
PAGE (10% gel) |
|
10 |
Es4 |
Esterase 4 |
Kidney |
CAE |
|
11 |
Es6 |
Esterase 6 |
Testis |
IEF (pH5–7) |
|
12 |
Es7 |
Esterase 7 |
Testis |
IEF (pH5–7) |
|
13 |
Es8 |
Esterase 8 |
Testis |
IEF (pH5–7) |
|
14 |
Es9 |
Esterase 9 |
Testis |
IEF (pH5–7) |
|
15 |
Es10 |
Esterase 10 |
Testis |
IEF (pH5–7) |
|
16 |
Es14 |
Sex-influenced esterase |
Plasma |
CAE |
|
17 |
Fh1 |
Fumarate hydratase 1 |
Kidney |
CAE |
|
18 |
Gc |
Group specific component |
Plasma |
PAGE (10% gel) |
|
19 |
Hbb |
Hemoglobin beta chain |
Red blood cells |
CAE |
|
20 |
Mup1 |
Major urinary protein 1 |
Urine |
PAGE (15% gel) |
|
21 |
Svp1 |
Seminal vesicle protein 1 |
Seminal vesicle fluid |
CAE |
|
Immunological marker |
||||
|
|
RT1 |
Histocompatibility 1 |
Red blood cells |
Hemagglutination |
|
CAE: Cellulose acetate membrane electrophoresis PAGE: Polyacrylamide gel electrophoresis IEF: Isoelectric focusing |
||||
Summary and Discussion
The results of this study can be summarized as follows:
|
1. |
Genes uniquely identifying each colony were found. |
|
2. |
Lower polymorphism observed in Jcl:Wistar was caused by caesarian section rederivation of the colony. |
|
3. |
Genetic distance suggested that Crj:SD and Jcl:SD are the closest genetically. It was also revealed that the relation between Iar:Wistar and Donryu was very close yet genetically different from SD and Wistar colonies. |
Although geneticists and breeders recognize the importance of genetic monitoring of closed colonies, this issue has not been discussed. The major reason is
TABLE 2 Gene Frequencies of 21 Biochemical Markers in Six Outbred Stocks
|
|
Locus |
Allele |
Jcl:Wistar |
Crj:Wistar |
Iar:Wistar |
Jcl:SD |
Crj:SD |
Donryu |
|
1 |
Acon1 |
a |
0.0000 |
0.0000 |
0.0000 |
0.9580* |
0.5670* |
0.0000 |
|
|
|
b |
1.0000 |
1.0000 |
1.0000 |
0.0420 |
0.4330 |
1.0000 |
|
2 |
Ahd2 |
b |
0.1830* |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
|
|
c |
0.8170 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
3 |
Ahdc |
a |
0.0000 |
1.0000 |
1.0000 |
1.0000 |
0.9020 |
1.0000 |
|
|
|
b |
1.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0980 |
0.0000 |
|
4 |
Akp1 |
a |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
|
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
5 |
Alp1 |
a |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
|
|
b |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
6 |
Amy1 |
a |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
0.6500 |
1.0000 |
|
|
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.3500* |
0.0000 |
|
7 |
Es1 |
a |
0.0000 |
0.0000 |
0.7000 |
0.1920 |
0.3750 |
1.0000 |
|
|
|
b |
1.0000 |
0.9670 |
0.3000 |
0.0000 |
0.1670 |
0.0000 |
|
|
|
c |
0.0000 |
0.0330 |
0.0000 |
0.8080 |
0.4580 |
0.0000 |
|
8 |
Es2 |
a |
1.0000 |
0.9170 |
0.0000 |
0.7670 |
0.3080 |
0.0000 |
|
|
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
|
|
c |
0.0000 |
0.0000 |
0.5420 |
0.1670 |
0.0000 |
0.5920 |
|
|
|
d |
0.0000 |
0.0830 |
0.4580 |
0.0670 |
0.6920 |
0.4080 |
|
9 |
Es3 |
a |
0.3330 |
0.9170 |
0.0000 |
0.2920 |
0.3920 |
0.9500 |
|
|
|
b |
0.0000 |
0.0000 |
0.4250 |
0.0000 |
0.0000 |
0.0000 |
|
|
|
c |
0.0000 |
0.0000 |
0.2330 |
0.4670 |
0.0000 |
0.0000 |
|
|
|
d |
0.6670 |
0.0830 |
0.3420 |
0.2420 |
0.6080 |
0.0500 |
|
10 |
Es4 |
a |
0.0000 |
0.0000 |
0.5420 |
0.1580 |
0.0000 |
0.0000 |
|
|
|
b |
1.0000 |
1.0000 |
0.4580 |
0.8420 |
1.0000 |
0.1420 |
|
|
|
c |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.8580* |
|
11 |
Es6 |
a |
1.0000 |
0.5520 |
1.0000 |
0.4330 |
0.6670 |
1.0000 |
|
|
|
b |
0.0000 |
0.4480 |
0.0000 |
0.5670 |
0.3330 |
0.0000 |
|
12 |
Es7 |
a |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
|
|
b |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
13 |
Es8 |
a |
0.0000 |
0.9170 |
0.4330 |
0.2330 |
0.7170 |
0.1330 |
|
|
|
b |
1.0000 |
0.0830 |
0.5670 |
0.7670 |
0.2830 |
0.8670 |
|
14 |
Es9 |
a |
1.0000 |
0.8330 |
0.5670 |
0.7650 |
0.2830 |
0.9150 |
|
|
|
c |
0.0000 |
0.1670 |
0.4330 |
0.23350 |
0.7170 |
0.0850 |
|
15 |
Es10 |
a |
1.0000 |
0.9170 |
0.5670 |
0.7830 |
0.2850 |
0.9170 |
|
|
|
b |
0.0000 |
0.0830 |
0.4330 |
0.2170 |
0.7150 |
0.0830 |
|
16 |
Es14 |
a |
1.0000 |
1.0000 |
0.7330 |
0.0000 |
0.5830 |
1.0000 |
|
|
|
b |
0.0000 |
0.0000 |
0.2670 |
1.0000 |
0.4170 |
0.0000 |
|
17 |
Fh1 |
a |
1.0000 |
0.0000 |
0.0000 |
0.0250 |
0.0000 |
1.0000 |
|
|
|
b |
0.0000 |
1.0000 |
1.0000 |
0.9750 |
1.0000 |
0.0000 |
|
18 |
Gc |
a |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
1.0000 |
|
|
|
b |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
0.0000 |
|
19 |
Hbb |
a |
1.0000 |
0.3170 |
0.7830 |
0.4250 |
0.0000 |
1.0000 |
|
|
|
b |
0.0000 |
0.6830 |
0.2170 |
0.5750 |
1.0000 |
0.0000 |
|
20 |
Mup1 |
a |
0.0000 |
0.0000 |
1.0000 |
0.0000 |
0.0350 |
0.9420 |
|
|
|
b |
1.0000 |
1.0000 |
0.0000 |
1.0000 |
0.9650 |
0.0580 |
|
21 |
Svp1 |
a |
1.0000 |
0.5670 |
1.0000 |
0.4000 |
0.7170 |
1.0000 |
|
|
|
b |
0.0000 |
0.4330 |
0.0000 |
0.6000 |
0.2830 |
0.0000 |
|
* Bold numbers denote stock-specific antigens. |
||||||||