Understanding the Introduction of Pathogens into Humans: Preventing Patient Zero: Proceedings of a Workshop (2026)
Chapter: 4 Forecasting, Surveillance, and Early Warning
4
Forecasting, Surveillance, and Early Warning
Highlights of Key Points Made by Individual Speakers*
- Applying principles of biology and ecology can iteratively constrain the risk space and inform more targeted application of resources for disease surveillance. (Han)
- Health of humans, animals, plants, and the environment are linked, and ecosystem changes affect them all. (Charles)
- Many issues in pathogen and disease surveillance can be overcome by using the power behind the One Health approach in artifical intelligence (AI). (Charles)
- Detection systems do not need to provide a representative picture of the population but need to investigate and have the ability to oversample high-risk areas. (Lipsitch)
- AI can play a transformative role in effective disease surveillance in resource-constrained countries. (Sengupta)
- Local input and leadership are important to create effective global health security solutions. (Charles, Han, Lipstitch, Sengupta)
* This list is the rapporteurs’ summary of points made by the individual speakers identified, and the statements have not been endorsed or verified by the National Academies of Sciences, Engineering, and Medicine. They are not intended to reflect a consensus among workshop participants.
The third session of the workshop was focused on the future applications of technology to enhance forecasting, disease surveillance, and early warning. Sumiko Mekaru, Epivoyant, LLC, began by describing the wide scope of this problem and encouraging the audience to consider the question of how this group can better anticipate risks of pathogen introduction in an evolving landscape.
PREDICTION AND FORECASTING
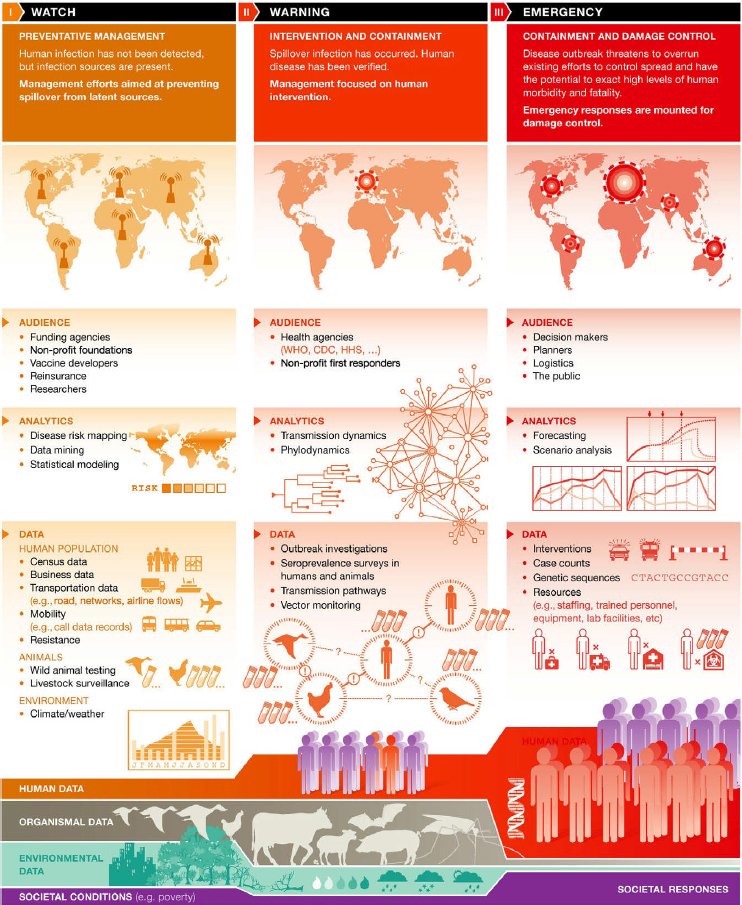
Barbara Han, Cary Institute of Ecosystem Studies, started the discussion by defining prediction and forecasting. She said that forecasting assumes that events occur as we move forward in time, and researchers can use information from the past to predict future occurrences. Forecasting can be done in circumstances where there are sufficient data on a particular system, host, or other point of interest. Forecasting is what modelers spend most time on during the emergency phase of an outbreak when attempting to contain or control damage (Figure 4-1).
Han explained that most work with data and subsequent models built to perform real-time forecasting happens in the emergency phase when working to control a pathogen (Figure 4-1) (Han and Drake, 2016). When the emergency is under control, there is a tendency to shift back into a “peace time” phase of research in which the pathogen is known and perhaps reliable diagnostics are available. Research can then focus on understanding what caused the pathogen to emerge, what elicited the first spillover, and what host species may have led to human exposure, she said. These foundational questions can only be answered if the type of pathogen and its location is known. In other words, the types of data required to make predictions include the population at risk, the organisms involved, and the environment in which those organisms are found. Comparatively little time is spent in this watchful phase if there is little information on the pathogen (Figure 4-1).
While a significant amount of data exists for priority pathogens, Han said, most locations are undersampled, most hosts are understudied, and most pathogens are undescribed. When thinking about early warning signals and early warning systems, Han noted that it is important to remember that prediction is quite difficult because of the number of unknowns that persist (Han and Drake, 2016). Some information can be drawn from the first principles of biology and ecology, such as making distinctions about whether the host has a particular receptor that could make it susceptible to a pathogen. Applying biological principles to iteratively constrain the risk space allows for a more targeted application of resources to conduct research and design action plans that inform surveillance.
Han noted that some exciting advances have come from work with filoviruses, Ebola virus, and SARS-CoV-2, for which a combined analysis
SOURCE: Presented by Barbara Han, January 15, 2024; Han and Drake, 2016.
of molecular structures and ecological traits and evolutionary principles of the host species were used to make specific predictions about which host species should be capable of becoming infected and could serve as reservoirs (Han et al., 2016; Lasso et al., 2025). Han described a project that aimed to better predict which bat species to sample in order to identify a reservoir species for filoviruses. The team analyzed biological and ecological traits
of the bats to accurately predict hotspots where potential seropositive bats might be, and which bat species had the highest risk of being a reservoir for filoviruses. This helped guide prioritization of sequencing and some field campaigns, Han said. In the priority bat species identified, researchers found new Ebola viruses, like Bombali virus discovered in Sierra Leone in 2018, and Sudan virus, which had an outbreak in 2022 in Uganda.
Han said that her team’s work in Southeast Asia continues to discover new filoviruses that have different sequences than previously detected in all countries where this work have been performed. Moving forward, she noted that it is essential to decide which filovirus species and bat species are of interest, where these can be found, and when they should be sampled. Han explained that experimental assays and data on ecology, land use, and livestock–human interactions are used to improve predictions. With the help of machine learning methods, some actionable decisions can be made about where to best invest funds for surveillance, she concluded.
AI-DRIVEN HEALTH SECURITY
Lauren Charles, Pacific Northwest National Laboratory (PNNL), introduced PNNL’s AI-driven One Health security program, which studies the health security of humans, society, animals, plants, and the environment. Charles explained that her motivation for creating this program stemmed from observing the limited collaboration between and within government and nongovernment entities during the COVID-19 outbreak. She found that data sharing and integration was impeded by the lack of data standardization. Further, the detection of threats and anomalies were delayed and insufficient coordination and cross-awareness led to inadequate responses to health threats, she added.
Charles also reflected on the current highly pathogenic avian influenza (HPAI) zoonotic outbreak, which has reached every continent and jumped to over 50 new animal species, including U.S. dairy cattle, which has led to infections in humans and cats. COVID-19 and HPAI are examples of ongoing problems in rapidly detecting and responding to outbreaks using current approaches, she said. Additionally, climate change and increased frequency of extreme weather events can disrupt food systems and increase the presence of new and emerging diseases, zoonosis, and even chronic illnesses, which represents a threat to One Health security.
PNNL’s AI-driven One Health security approach recognizes that health and security of humans, society, animals, plants, and the environment are all intimately connected through complex interrelationships. Charles noted that AI methods that have been developed over the past decade can detangle these intricacies. By applying AI algorithms, she added, researchers can improve early warning systems and better monitor, predict, detect, control, and mitigate health security threats, whether natural, intentional, or accidental.
Most current approaches for biosurveillance are siloed, Charles continued, where each domain is focused only on identifying threats within a specific area or species of concern, with minimal cross-communication or data sharing. Additionally, often only one type of data, such as case counts, is used during analysis. Unfortunately, these approaches offer limited ability to detect anomalies and fully inform situational awareness, Charles explained, which can lead to significant gaps in disease prediction and risk assessment. Approaching this problem with a One Health lens shows that, because the health of humans, animals, plants, and the environment are all linked, each will be affected in some way by ecosystem change. Therefore, animals, plants, and the environment can serve as early warning sensors for human health threats, informing deployment of countermeasures even before humans are affected, Charles claimed.
Charles noted that her research team has been able to overcome many challenges in disease surveillance by applying the One Health approach using AI tools. Charles added that her team is collecting heterogeneous datasets across disciplines, applying new multimodal data harmonization techniques, and applying AI to derive actionable insights, uncover patterns, and forecast potential direct and indirect health threats. Charles said they have been successful because they are focused on building trusted partnerships with data providers and federal sponsors across all sectors. This is an essential part of the One Health approach, she said.
Finally, Charles highlighted some of the tools that PNNL has built to demonstrate the applications of the AI-driven One Health security approach for situational awareness, early warning risk, and prediction (Pacific Northwest National Laboratories, n.d.):
- Biofeeds is a generalized data harvester that pulls in open-source news and reports from around the world and runs analytics to help users identify new, ongoing, or potential events that may cause harm to humans, animals, or plants, and extracts characteristics and impacts of those events (Baker et al., 2018).
- The MedINT platform, or medical intelligence, focuses on medical operations in early warning from a top-down perspective, taking an all-source approach for event detection that includes open-source news, scientific literature, animal health reports, and severe weather events to gain global situational awareness of potential national health threats, which includes potential disease introductions. It uses a type of syndromic surveillance to identify baselines rising and anomalous phenotypes within medical records.
- The One Health (OH)-TREADS platform enables individual states to effectively share health data between agencies. Sophisticated data sharing and cybersecurity data harmonization techniques enable insights drawn across human society, animal, and pathogen
- Zoonotic Spillover Risk (ZooRisk) uses data from open-source news platforms and identifies zoonotic diseases reported in humans or animals and then extracts pertinent information to run simulation models of the risk that these diseases present to humans based on a variety of factors from census data to special events like state fairs.
- Machine Learning for Disease, or ML4Dz, uses thousands of indicators to help overcome biases and gaps in infectious disease data and enable the prediction of disease up to 1 year in the future, even in locations with few disease reports.
- National Wildlife Disease Database takes both a top-down and a bottom-up approach for early warning and situational awareness to better detect, prevent, and mitigate threats to wildlife health before these threats potentially cause an outbreak in humans.
vector data. In addition to applying AI for situational awareness and anomaly detection, it also creates risk maps of current disease threats and forecasts, even when there is a lack of data and therefore a higher risk of bias in the available data.
DATA INTEGRATION AT EARLY STAGES OF OUTBREAKS
The next panel member, Marc Lipsitch, Harvard T.H. Chan School of Public Health, began by noting that people often understand the need for disease surveillance but do not always see it as a set of interrelated activities that need to be done in specialized ways for different purposes (Lipsitch et al., 2024). He used the metaphor of the Swiss Army knife, with each part having a distinct purpose. Similarly, the disease surveillance systems that are designed to detect new unexpected threats are different from those that are designed to focus on individual threat assessment and treatment or characterizing severity or burden of disease. Disease detection systems do not need to provide a representative picture of the entire population, he said, but need to investigate high-risk areas and have the ability to oversample specific areas that are most likely to be positive. However, he added that if the goal is to understand disease burden, such as during the COVID-19 pandemic, a representative sample is needed.
Lipsitch said that there is always immense uncertainty about what is happening now and what may happen in the future. For instance, in January 2020 a news report declared that influenza was a bigger problem than SARSCoV-2. While this was true at the time of the report, SARS-CoV-2 rapidly emerged as the greater challenge. Identifying a real, pressing threat is challenging, he said. For instance, during the first months of the H1N1 pandemic in 2009, studies demonstrated significant uncertainty regarding the case fatality for this novel viral strain that caused an influenza pandemic (Garske
et al., 2009; Wilson and Baker, 2009). Plans for pandemic preparedness and response are often made based on severity, Lipsitch said, but severity may not be known for many months, which makes it challenging to choose the most appropriate plan at the time of the ongoing disease outbreak.
Lipsitch described the value of model-based thinking, in which disease transmission modelers think quantitatively about disease transmission. Good modelers, he added, understand where data come from and what aspects of data may be missing and biased, leading to understanding what questions to ask to address some of the key components of risk. Finally, experience in the United Kingdom during the COVID-19 pandemic has shown the value of integrating infectious disease modeling and the ideas of the practitioners into the design of disease surveillance systems.
An example of model-based thinking in action is the Center for Forecasting and Outbreak Analytics (CFA) at the Centers for Disease Control and Prevention (CDC), which has developed a model-based, qualitative risk assessment. This tool is updated frequently and contains detailed risk assessments for different segments of the population (CDC, 2025). It also differentiates the likelihood and the effect of infection in its risk assessment. When the risk is considered low, it can be caused by a low probability of becoming infected, a small effect of the infection, or both. When using the word risk, he added, it is important that the meaning is specified in context. He said that it is also essential that public health officials are open to shifting their guidance as new information becomes available.
An example of integration, Lipsitch said, is interdisciplinarity. He asserted that there is a need for multiple types of evidence to inform a response to a potential or actual pandemic (Lipsitch, 2020). In times of intense uncertainty, Lipsitch added, experts tend to stay within their disciplines instead of seeking collaboration. Lipsitch argued that this is the wrong response, though it is an understandable sociological action.
During a public health threat, decisions must be made rapidly and informed by multiple disciplines. The importance of interdiciplinarity is demonstrated in the example of research on masks and airborne transmission of COVID-19. At the beginning of the pandemic, some important publications were available detailing the physics of airborne transmission and the epidemiologic evidence of the role that masks could play in reducing transmission of other coronaviruses. Each of these publications provided evidence to inform decisions at the pace required for a public health emergency.
Lipsitch concluded by describing the importance of being able to adjust opinions and recommendations based on the most recent and relevant data, which change rapidly early on in an outbreak response cycle. However, once an urgent decision has been made, it is easy to either overrespond or have a response that outlasts its usefulness, he said. It is
essential to have opportunities to change policy in a structured way. Analytics, disease surveillance, and modeling can be strengthened so a credible evidence base can be provided when recommendations are changed. He posited that quickly made decisions should have an expiration date and should be revised regularly. Releasing data on a regular schedule can help the public understand that facts change and to expect regular updates without being alarmed when new data emerge. This process should start in “peace time,” Lipsitch said, because public expectations take time to develop. Honesty from science experts may help make communication more effective, he said.
AI GOVERNANCE AND GLOBAL INCLUSION
Aparupa Sengupta, Nuclear Threat Initiative (NTI), presented her experiences from working in various sectors and also shared comments prepared by Sarah Carter, Science Policy Consulting LLC.
Sengupta began by discussing the application of AI tools to enhance disease surveillance and forecasting. AI tools present a powerful opportunity to improve public health, she said, especially in regions with limited resources. She also addressed the international context of AI governance and the importance of global inclusivity to ensure that the development and deployment of AI technologies benefit all nations, not just nations where technology advancement is currently concentrated.
Regions in the Global South often face resource constraints, said Sengupta, which can hinder effective disease surveillance and forecasting. She posited that emerging technologies like AI can play a transformative role. AI algorithms can analyze diverse data sources, such as news reports, for forecasting patterns in disease surveillance. Social media tools such as Health Map and Epi Wash can also identify unusual symptom clusters or frequent disease mentions, she said. Machine learning models can identify patterns in historical disease and climate data and forecast outbreaks with increasing accuracy. Accuracy and speed are required when there is a disease outbreak to allow authorities to allocate resources appropriately and implement preventative measures. AI can also facilitate real-time monitoring by automating data collection and analysis from various sources, including electronic health records, mobile health apps, and variable sensors. She explained that AI can enable identification of trends and risk factors in real time. Further, medical image analysis by AI may help to quickly and accurately diagnose diseases even when trained specialists are limited, which is often the case in low- and middle-income countries.
Sengupta mentioned that several tools are publicly available and can be used for disease surveillance and forecasting. However, to gain a
comprehensive understanding and maximize benefits of this technology, it is essential to acknowledge its challenges and vulnerabilities. Data scarcity and quality, especially in the Global South, make data less reliable and can reduce the effectiveness of these AI tools, she said. Infrastructure limitations are another significant challenge, as computational resources and Internet access remain limited in some areas, she explained.
There are also ethical issues regarding data privacy algorithm bias, Sengupta added. The way algorithms have been developed and the types of data they are trained on will determine their output. There is also an equity issue in terms of access to technology, she said, especially regarding the Global South or resource-limited countries. Further, there is a risk for biological threats when AI tools for biosurveillance identify harmful new viral variants. If a malicious actor obtains such information, they have an opportunity to turn the new pathogen into a biological weapon. Further, bad actors could use generative AI to develop ideas on how to target vulnerable populations, agriculture, and infrastructure, she said. Another risk is that these frontier AI models may also contribute to the generation of misinformation and disinformation. Drawing from her experience and work in the Global South, Sengupta noted that this is a major problem in many countries.
Sengupta then discussed the international context of AI governance and the importance of global inclusivity. She asserted that it is crucial to ensure that the Global South has a voice in shaping the future of AI. Given the global nature of disease threats, she stated, all voices must be heard to create effective global health security solutions. Global stakeholders should support AI research and development in the Global South and promote development of culturally relevant and contextually appropriate AI solutions, as these solutions are based on the local conditions, infrastructure, availability of skilled professional, and other local data sources. As such, she cautioned about imposing AI governance models developed in the Global North on the Global South because the social context and needs are significantly different.
Sengupta presented some examples and ongoing work to address these vulnerabilities and challenges. The Africa CDC Pathogen Genomic Initiative 2.0 promotes real-time data sharing for informed public health actions. The increased availability of affordable portable metagenomic sequencing tools is facilitating their access in more remote and low-resource settings. Africa CDC is also working to build an agile and responsive AI-enabled biosurveillance network. Sarah Carter has also worked with the team at NTI to develop safeguards for AI-enabled biological design tools, added Sengupta. Much of the modeling of these tools curated for biomedical users or predictive disease modeling could present challenges if the risk or vulnerabilities of the design are not assessed.
The team has made progress is establishing guardrails and best practices to mitigate vulnerabilities as much as possible. The AI-bio Forum is an international platform where experts convene with diverse stakeholders from the Global North and South to discuss biosecurity risk posed by the rapid advancement of AI technologies (NTI, 2024). Sengupta explained that a goal of the forum is to represent perspectives on regional vulnerabilities. This type of work can foster greater international collaboration and facilitate international health security with the goal of mitigating disease transmission, she said. Sengupta closed her talk by stating that more collaborative efforts for responsible and equitable development of AI are under way.
DISCUSSION
Turn Data into Action
Mekaru began the discussion by asking how the vast amount of data being generated and collected can be translated into action. Charles replied that with currently available tools, especially AI, it has become easier to combine and model data to get results. However, it is important to consider what data to combine, how to combine them, and the appropriate methods to analyze the data, she said. Further, missing data, data biases, differences in collection protocols, uncertainty propagation, and other factors should be considered. Every dataset has uncertainty, she said, which can be compounded when combining multiple datasets. Finally, she said, when working across disciplines, subject-matter experts for each data type need to be involved. Han further emphasized the need for interdisciplinary experts, as it improves data accuracy but also unlocks additional answers to downstream questions that may not be considered otherwise.
Online participants asked whether current models and technologies would have been able to identify the points of entry—where and when—that cattle would have been exposed to and infected with H5N1 if all data were available. Han replied that if all data were available, it is possible that accurate predictions could have been made. However, she added, it does not necessarily inform the best action to take. In addition to being able to make the prediction, it is equally important to know what to do with it, she said. Charles added that models are never 100 percent correct, but they can point you in the right direction to investigate whether precautions are needed. She also said that citizen science is an area that could be considered to fill some data gaps.
Collaboration
In discussing collaboration, Han said that answering interdisciplinary questions requires different data streams and experts who connect data and inform a model to create predictions that are actionable and accurate. Such an approach takes time and requires a common language and support, she added. Sengupta reflected that the AI-bio Forum has shown that bringing people together results in different perspectives being presented, which is essential for harmonized guidelines and policy decisions. Breaking large groups into smaller working groups can be helpful in identifying solutions more quickly, she said. Charles added that collaborations lead to bigger networks and that trust is easier to obtain when there are previously built relationships. Collaboration enables better analytics and technological advancements and may be required for access to certain data. When building collaboration, it is also important to consider how to incentivize and clarify rules and processes on data sharing, she said.
Unconventional Data Sources
Mekaru asked the panel about the advantages of using unconventional data sources, such as wastewater surveillance and the use of social media and satellite imaging. Charles said that there are nuances to each data source, and gaps left by unconventional sources can be filled with data from conventional sources. Social- and health-related data, such as gross domestic product or child morbidity rates, can serve as proxy data to fill gaps in disease forecasting systems. These data may facilitate more accurate predictions of impact even if the precise connection to a pathogen is unclear. Han added that unconventional data sources are useful when information is needed for rapid decision making. She added that it is also important to establish baseline data for every measure to establish situational awareness of a healthy system.
Equitable Access and International Collaboration
Mekaru said that equitable access or international collaboration are not only questions of morality but also provide concrete benefits. Han agreed and said that no good science can be done without engaging with people who are knowledgeable about the system, which includes local experts. Sengupta added that disease outbreaks begin locally, and thus each outbreak exists within a unique context. Localized expertise is critical to conducting good science and developing sound policy that supports a robust global health security network.
Lipsitch provided an example of the issue. Laboratory capacity has expanded in the southern African region through investments that were partially domestic and partially from the Global North. This capacity made it possible to identify the Omicron variant of SARS-CoV-2 and issue a global alert; however, the global alerts resulted in travel bans to the countries that made the identifications. These disincentives may limit reporting of important disease or pathogen findings in the future. Sengupta added that if basic transparency and trust is lacking, then no progress can be made. Therefore, it is important to involve local and regional experts in the discussion, starting from data modeling to algorithm development, deployment of tools, and policy discussions, she said.











