Firepower in the Lab: Automation in the Fight Against Infectious Diseases and Bioterrorism (2001)
Chapter: Laboratory Firepower for AIDS Research
6
Laboratory Firepower for AIDS Research
Scott P. Layne and Tony J. Beugelsdijk
INTRODUCTION
HIV poses enormous challenges from a variety of directions. In less than three decades, it has grown from an unknown pathogen to a pandemic disease, chronically infecting more than 50 million people and killing nearly 3 million people each year. By the beginning of twenty-first century, an estimated 1 percent of the world's population will be afflicted, and end-stage AIDS will rank as one of the top five causes of death by infectious disease (Murray and Lopez, 1997). In response to this catastrophe, pharmaceutical companies are manufacturing new antiviral drugs (e.g., reverse transcriptase and protease inhibitors) that extend peoples' lives and investigators are examining chemokine receptors (e.g., CCR and CXCR) and cytotoxic cellular epitopes that offer promising leads for therapeutics and vaccine development (Baggiolini, 1998; Heilman and Baltimore, 1998). Despite such rapid progress, however, the fact remains that individual investigators tend to shy away from important problems dealing with enormous “experimental spaces” simply because they lack the necessary tools to move ahead.
The current models of discovery and follow-up are based on investigator-initiated research and relatively small-scale efforts, but in waging the war on AIDS the research community should also consider how intermediate-scale research infrastructures (first referred to as “collaboratories” by William A. Wulf) could feasibly contribute (National Research Council, 1993). Basic as well as clinical investigators consequently should ask three questions about the potential roles for such
collaboratories. First, what activities in AIDS research (see Box 6.1) require enormous inventories of data and information? Second, what categories of laboratory-based tests (see Figure 6.1) must be carried out to create such inventories? Third, what technologies are available for creating high-throughput research facilities that are cost effective, reliable, and flexible? To move forward with collaboratory-based efforts for AIDS research, there must be some form of consensus among investigators and administrators. After all, scientific and medical research requirements must drive the demand for high-throughput resources, not vice versa.
If sufficient agreement does indeed exist, a critical number of available technologies and scientific disciplines could certainly be brought together to level the playing field against the growing threat of HIV infection and AIDS mortality. The available tools would include reliable laboratory tests (methods), automation and robotic equipment (hardware), object-oriented programming languages (software), relational databases (informatics), shipping services (virtual warehousing), and Internet providers (communications). This paper sets forth, with such tools in hand, a blueprint for creating flexible laboratory and informatics facilities that could be operational within 2 to 3 years for basic science and a variety of clinical trials and public health efforts. As described later, important roles for such laboratories could include development of HIV vaccines and optimization of therapeutic regimens by investigators on national and international scales.
|
BOX 6.1 Important Problems in Basic and Clinical AIDS Research that Could Benefit from Newer Models of High-Throughput Laboratory Research.
|

FIGURE 6.1 Throughout the world, HIV-infected individuals harbor an extremely broad array of wild-type viruses, and AIDS investigators have limited systematic information on these pathogens. The lack of basic knowledge of how viral reproduction, genetic mutation, and immune selection are interrelated raises the issue of whether comprehensive surveys should be initiated. Such surveys would build inventories of epidemiologic and laboratory-based data and seek correlations (if any) between phenotypic expressions, genotypic variations, and disease progression. The figure shows typical measures that apply to each category of data. Ultimately, it may be possible to identify the “right” assortments of HIV genotypes and phenotypes for vaccine development. Logical considerations would include the most prevalent and representative strains, but there is no clear consensus on this issue at present. An inventory of systematic information could help.
CURRENT LIMITATIONS
Unprecedented digital technologies are now at the fingertips of scientific and medical investigators in many geographic locations. To illustrate this point, consider the magnitude of three representative benchmarks from supercomputing, information storage, and worldwide communications. At many advanced computer facilities, numerical performances are surpassing ~1012 floating point operations per second (teraflop speeds), database storage facilities are exceeding ~1015 bits per site (petabit capacities), and Internet communication rates are achieving ~109 bits per second (gigahertz bandwidths). Despite such fantastic digital firepower, many important AIDS research efforts currently are limited by sheer physical firepower. The biggest rate-limiting step is the ability to perform vast numbers of tests in the laboratory (Layne and Beugelsdijk, 1998).
Most laboratory experiments in AIDS research are performed by human hands, usually in combination with sprinklings of labor-saving devices and small islands of automation. This semimanual approach may certainly lead to insights and breakthroughs, but such laboratory work is often repetitive and exceedingly tedious. Even with an army of post-doctoral students and laboratory technicians, human hands still remain the limiting factor in generating the vast quantities of raw information required for solving complicated problems. Fortunately, rather impressive technological advancements have been made in recent years, and so the time is now right for overcoming such manpower-related obstacles.
Virologists, immunologists, and molecular biologists have developed a variety of laboratory-based assays that are reproducible and readily adaptable to large-scale efforts. Engineers have developed innovative automation and robotics technologies that are capable of skyrocketing the number and variety of laboratory experiments. Computer scientists have developed Internet programming languages and database management systems that are providing the basic building blocks for improved environments in scientific collaborations. And physicists (driven by the need to share gargantuan amounts of data generated in high-energy particle physics experiments at a few large accelerator facilities) have developed the World Wide Web. Today, these developments are literally transforming the ways in which scientific collaboration and information distribution are taking place (Lanier et al., 1997). As described below, the integration and refinement of these capabilities hold significant promise for accelerating AIDS research in basic science, clinical trials, and public health efforts throughout the world. The objective is to create a high-
throughput laboratory that is practical, flexible, and easy to use from any geographic location.
BATCH SCIENCE
The Internet can be used by laboratory-based investigators in either of two ways—for handling a series of real-time or nonreal-time operations. Real-time operations require specialized communications protocols that are redundant and failsafe. An actual illustration of real-time manipulations is access to a multiuser scanning electron microscope from a distance. The user operates the microscope's controls in one location, and the sample is manipulated and viewed while residing in another place (Chumbley et al., 1995). In general, few specialized applications in science and medicine require real-time operations from afar. Moreover, the Internet's hard wiring and protocols are not ready for handling large volumes of real-time manipulations every day, which will require much larger bandwidths (Germain, 1996).
On the other hand, nonreal-time operations do not require specialized communications protocols that are redundant and failsafe. Most scientific and medical research efforts can be supported by nonreal-time operations that are scripted with flexible software tools—an approach that will be referred to as batch science via the Internet (see Figure 6.2). Batch science machines would perform the manipulative work of hundreds of humans, serve as programmable laboratory technicians, and help AIDS investigators tackle certain types of big problems (Box 6.1).
A feasible illustration of batch science would be the undertaking of major AIDS vaccine trials (perhaps in the near future) with the assistance of an automated reference laboratory. In this example, investigators would collect numerous clinical samples from vaccinees before and after receiving the series of immunizations (i.e., the efficacy cohort) and, in a parallel effort, from HIV-infected individuals in close geographic proximity but not receiving such shots (i.e., the carrier cohort taking other forms of therapies). Investigators would then use flexible software tools to design and script the necessary laboratory tests while residing at their home bases. The available “toolbox” of laboratory tests would include full assortments of viral infectivity, molecular sequencing, and cytotoxic T lymphocyte assays (see Box 6.2). Next, investigators would airfreight the frozen samples to their collaborating facility (in another geographic location) and within just a few days the desired assays would be set up and carried out by high-throughput automation in accordance with the digital “assay scripts” that arrived via the Internet. Upon completing the necessary tests, data from the various assays would be electronically deposited into the trial's supporting database, and collaborating teams of investigators
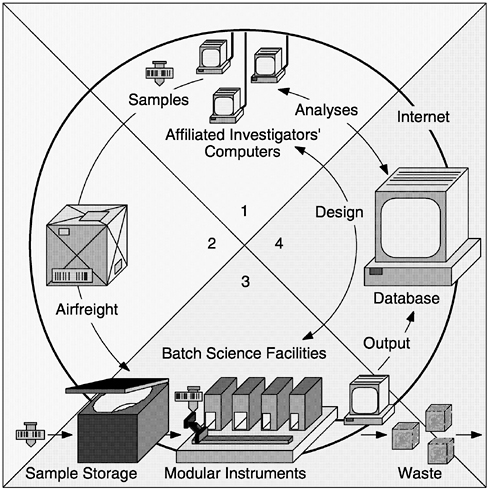
FIGURE 6.2 Batch science via the Internet would enable investigators (at any geographic location) to submit sets of biological samples and automated-instrument instructions as coordinated packets. This approach is distinct from virtual science via the Internet, where experiments are controlled from afar in real time. (1) Scientists located anywhere in the world design experiments in cooperation with a particular batch science laboratory. (2) Shippers deliver packages containing barcoded specimens or reagents. (3) Batch science laboratories house flexible, modular, and scalable instruments. (4) Database facilities maintain permanent records and provide software for analyzing and managing data. The flow of materials and information through the batch operating system is summarized below. The sequence of steps is well suited to most basic science, clinical trials, and public health efforts:
-
Investigators download Process Control Tools (PCTs) from the Internet.
-
Investigators design assay scripts at home base.
-
Investigators collect, package, and bar code numerous samples.
-
Investigators document background information.
-
Investigators e-mail assay scripts via the Internet.
-
Samples and special materials are shipped to the laboratory facility.
-
Laboratory receives/logs the bar-coded samples and digital assay scripts.
-
Laboratory holds the samples in short-term frozen storage.
-
Laboratory pairs the samples with the assay scripts.
-
Laboratory provides common supplies and reagents.
-
Laboratory performs assays in a batch science mode.
-
Laboratory disposes of used specimens and waste materials.
-
Laboratory archives certain specimens as ordered by investigators.
-
Laboratory forwards raw data to the database.
-
Investigators analyze their own data with additional sets of PCTs.
-
Investigators link their data to other relevant information.
-
Investigators assign data-use privileges.
-
The investigator-airfreight-laboratory-database cycle continues.
-
Organized digital records amass over time.
would use this information to address the following key questions as the effort moves forward (Anderson et al., 1997):
-
What are the vaccinees' baseline immune parameters?
-
What percentage of vaccinees respond to immunization?
-
What are the humoral and cellular responses after each dose?
-
Does the vaccine demonstrate protection or efficacy?
-
How broad are the responses against the collected panel of carrier viruses?
-
Are there any significant isolate-specific failures?
-
How long does protective immunity persist?
When compared to laboratory procedures performed by human technicians, the quality assurance team would further observe that batch science methodologies are significantly faster, offer larger sample sizes, and exhibit greater reproducibility—all of which would translate into far better clinical trials for AIDS vaccines.
To carry out the various procedures illustrated above, AIDS investigators would use a suite of Process Control Tools (PCTs) to program, coordinate, and track scientific procedures at every step. For instance, PCTs would be used to assign bar codes to samples, to script individualized assay protocols, to analyze raw data, and to create relational links with associated information. Also, at every step along the way private or
sensitive information would be protected by any number of accepted encryption and authentication methods. This would also be handled by the PCTs (see Box 6.3).
AVAILABLE TECHNOLOGIES
Modular automation and robotics hardware are now available for conducting practically every necessary task in AIDS research—such as bar coding, liquid handling, centrifuging, incubating, sequencing, immunostaining, scintillating, image capturing, and flow cytometer scoring (Garner, 1997). In addition, most major manufacturers of high-throughput laboratory equipment also sell software systems for interconnecting and operating their own products. However, the problem is that such proprietary systems work only so long as customers use equipment from a single company, which is rather unusual in many laboratory settings. Thus, a growing frustration for investigators is that it takes an inordinate amount of time to write new software that integrates components from various manufacturers into one effective machine.
To alleviate such problems, the American Society for Testing and Materials (ASTM) has recently adopted the Laboratory Equipment Control Interface Specification, or LECIS (ASTM, 1998). The new standard will enable manufacturers to create flexible software systems for interconnecting and operating practically any type of compliant hardware, and such innovations will soon lead to high-throughput laboratory systems that exhibit many compatibilities and conveniences found in personal computers. As indicated by the lowermost arrows in Figure 6.3, LECIS defines all the necessary commands between Standard Laboratory Modules (SLMs) and one Task Sequence Controller (TSC). From an investigator 's perspective, SLMs are programmable finite-state machines capable of performing various customized tasks in laboratory protocols. Suppose the first few tasks in a viral infectivity assay (for the vaccine trial mentioned above) are to thaw the sample tubes, open the tubes, and then add fresh culture media. In this example, a liquid-handling SLM would perform the following set of tasks under the overall command of the TSC:
SLM ready → input capped/frozen tube → inspect → warm to 37ºC → uncap → add liquid → recap → vortex → verify weight → output capped/reconstituted tube → SLM ready
In batch science machines, each SLM is dedicated to performing particular parts of laboratory tasks (e.g., moving, measuring, mixing, reacting, culturing) that fall within its physical constraints (e.g., volumes,
|
BOX 6.3 Process Control Tools (PCTs) to Program, Coordinate, and Track Scientific Procedures. PCTs would be fashioned to connect investigators from any geographic location to batch science machines and their associated database facilities. PCTs would be designed to permit maximum flexibility and control over experiments by remote investigators, just as though they were employing laboratory technicians to carry out their assay protocols. PCTs would also be designed to interface easily with web browsers (such as the Microsoft Explorer, Netscape Navigator, and Sun Hotjava) and data analysis software packages that are commercially available. To enable just about any type of research activity, one can envision a collection of seven basic PCTs as follows:
|

FIGURE 6.3 Batch science machines would operate on a three-tier hierarchy. At the lowest level, modules possess Standard Laboratory Module (SLM) controllers that drive components such as actuators, detectors, and servomotors and coordinate their internal electromechanical activities. Such events are contained wholly within each module (i.e., modules are blind to the existence of one another), which gives rise to independently working SLMs. At the intermediate level, Task Sequence Controllers (TSCs) use tools from operations research to govern intricate flows of supplies and samples through the entire machine. Various standardized commands and feedback signals work to carry out complete laboratory procedures, leading to flexible and programmable assay scripts. At the highest level, PCTs would enable AIDS investigators to carry out a spectrum of important activities via the Internet.
g forces, temperatures) and is also capable of performing these tasks in any arbitrary order (Beugelsdijk, 1989). In theoretical terms there is no upper limit to the number of SLMs per machine or to their physical size. In practical terms we envision that high-throughput machines for AIDS research could be constructed from a collection of merely 10 to 20 different SLMs that cover a full range of tasks. Analogous to computer architecture, duplicate modules can be used to add parallel-processing capabilities to machines, and expansion slots can be used to incorporate new capabilities or technologies. It is also feasible to build batch science machines based on conventional macroscale technologies (e.g., pipettes, test tubes, 96-well plates), newer nanoscale technologies (e.g., microchannels, microchambers, microarrays), or modular combinations of both (Persidis, 1998). Batch science machines thus offer unprecedented capabilities to carry out high-throughput research in a highly flexible operating environment. It now becomes a question primarily of the bigger strategies and directions that AIDS investigators wish to pursue.
THREE ESSENTIAL ASSAYS
Basic science, clinical trials, and public health efforts all rely on a laboratory-based toolbox that consists of viral infectivity, molecular sequencing, and cytotoxic cellular assays (Burton, 1997; Moxon, 1997; Bangham and Phillips, 1997). These three assays are then customized and optimized to meet the particular needs at hand (Box 6.2). Instrument designers would thus consider the most advantageous way to build a small assortment of batch science machines that work together as seamless units (Pollard, 1997). The bottom line is that small clusters of high-throughput machines would be flexible enough to cover the range of tasks that most AIDS investigators demand.
Traditionally, viral infectivity assays have been used in basic research, but more recently they have found clinical utility in the area of drug susceptibility tests (Martinez-Picado and D'Aquila, 1998). Because HIV mutates spontaneously and tolerates any number of amino acid mutations, it can readily escape the inhibitory pressures of reverse transcriptase and protease inhibitors. Consequently, the majority of HIV-infected individuals who take highly active antiretroviral therapies (HAART) are at risk for selecting multidrug-resistant isolates that predominate with time. In clinical practice these complications are first recognized by increasing HIV RNA levels and decreasing CD4+ cell counts, but, if left untreated, they can also proceed to accelerated disease progression (Hecht et al., 1998). Because multidrug-resistant isolates are becoming more common, there are growing demands for drug susceptibility tests (e.g., phenotypic assays) that provide useful information for clinical decision making. How-
ever, there are now 11 licensed antiretroviral drugs, and, potentially, each one must be evaluated over a range of concentrations in order to calculate the concentration that inhibits HIV growth by 50 percent (IC50), 90 percent (IC90), or both (Hirsch et al., 1998). The overall complexity and expense of selecting optimal therapies reinforce the need for high-throughput laboratory resources that are highly flexible and that take advantage of economies of scale.
Guiding clinical decisions are just one important use for viral infectivity assays. Others are listed in Box 6.2. Practically any design of infectivity assay would be carried out by three batch science machines that work in series. The first machine would input bar-coded samples, dispense cell cultures (e.g., lymphocytes, monocytes), add customized reagents (e.g., immunoglobulins, antivirals, chemokines), add viral inocula, and output bar-coded plates. The second machine would incubate incoming plates, replenish culture media at specified intervals, and output fully cultured plates. The third machine would score the incubated wells (e.g., by colorimetry, image analysis, flow cytometry) and then send the raw data to the information storage system. From the user's standpoint, all three viral infectivity machines would work together while following the commands of assay scripts (Figure 6.2). In addition, the first and second machines would be capable of performing all the necessary tasks for making recombinant viruses and growing infectious viral stocks.
Molecular sequencing assays have also moved beyond the realm of basic research and into the realms of clinical care and public health efforts. One such application pertains to detecting point mutations in reverse transcriptase and protease enzymes that confer various levels of resistance to one or more antiretroviral drugs. Depending on the choice of laboratory techniques, these genotypic assays are capable of yielding either full-length or partial-length sequences, but, in either instance the amino acid data offer only partial answers (Martinez-Picado and D'Aquila, 1998). In many situations, resistance is conferred not only by the point mutations themselves but also by complex structural and compensatory interactions throughout these viral proteins. To gain more accurate assessments, genotypic assays must therefore be interpreted in conjunction with the phenotypic assays described above (Hecht et al., 1998). This need to perform two categories of laboratory tests and organize multiple sets of data is well suited to batch science methodologies.
Another important application for molecular sequencing assays pertains to identifying the “right” assortment of viral sequences for inclusion as immunogens in future AIDS vaccine trials. One logical approach would consider sequences that represent the most prevalent genotypes from certain geographic regions, but there is no clear consensus on the issue at this time. A comprehensive survey of viral genotypes could certainly help with
this identification process; however, such an undertaking will require enormous numbers of epidemiologic samples and sequencing assays ( Figure 6.1). AIDS investigators could feasibly conduct such large-scale surveys with flexible PCTs (Box 6.3) that work in conjunction with high-throughput viral sequencing facilities.
Practically any design of molecular sequencing assay would be carried out by three batch science machines. The first would complete sample preparation and polymerase chain reaction (PCR)-amplification steps on cell-free viruses that contain RNA-based cores and cell-associated viruses that harbor DNA-based messages. The second and third machines would work in parallel and house modules for short-length (chip array and mass-spectrometer-based) or long-length (electrophoresis and microcapillary based) genomic determinations, respectively (Garner, 1997). All three molecular sequencing machines would work together in accordance with assay scripts.
In general, cytotoxic T lymphocyte (CTL) assays have been the most difficult to utilize in basic research and clinical management situations. One problem has been the lack of laboratory methods for direct visualization and quantification of antigen-specific cytotoxic T cells (Callan et al., 1998). Another has been the supply of sufficient quantities of HLA-restricted T cells for large-scale research efforts (Schwartz, 1998). But recently a number of innovative laboratory methods have been developed for reducing these problems, enabling AIDS investigators and clinicians to consider how batch science facilities could enhance their work.
Most CTL assays are set up to measure the “percent specific lysis” by chromium release, which is generally scored at one time point after mixing various ratios of effector cells (E) with target cells (T). Although such E:T assays are often used to detect specific CTL activities in vitro, they are limited to providing only static measures of cell killing (Pantaleo et al., 1997; Rosenberg et al., 1997). On the other hand, scoring CTL assays over a series of time points and utilizing new computer-based methods of data analysis would make it possible to provide kinetic measures of cell killing (Spouge and Layne, 1999). In such experiments the control arm would measure HIV reproductive statistics (e.g., viral burst sizes and cycle times) in the absence of effector CTL, while the corresponding experimental arm would measure reproductive statistics in the presence of effector CTL, with all other conditions kept similar to controls. If the cytotoxic cells have specific activity, they would perturb one or more reproductive statistics (e.g., smaller burst sizes, longer cycle times) in a concentration-dependent pattern. If the cytotoxic cells have no specific activity, reproductive statistics would demonstrate only random fluctuations or possibly patterns attributable to nonspecific cellular toxicity.
Intact cellular immunity is now deemed important for survival in
HIV-infected individuals (Pantaleo et al., 1997; Rosenberg et al., 1997). Since cellular immunity is inherently a kinetic process (e.g., cell killing rates versus viral reproduction rates), it is conceivable that time series CTL assays would offer more relevant assessments of cellular immunity than static E:T assays. Nevertheless, CTL assays that are set up for static (one time point) or kinetic (multiple time points) measures of cell killing would be performed by two batch science machines that work in series. The first machine would complete sample preparation and set up the desired CTL assays. The second would score the CTL assays by scintillation or flow cytometry over the desired number of time points, with both working together according to assay scripts.
In many areas of AIDS research it is often difficult to compare the results from one investigator's laboratory with those of another. Different lots of reagents, minor changes in experimental protocols, and inconsistent cell culture conditions can all lead to variable outcomes (Layne et al., 1992). On top of this, viral infectivity, molecular sequencing, and cytotoxic cellular assays display an inherent level of noise—often necessitating many replicate measurements and painstaking quality controls to guarantee reproducibility. Batch science facilities could help AIDS investigators overcome such difficulties in two meaningful ways. First, the programmable machines would have the necessary flexibility and firepower to evaluate which assay formats are the most reliable and reproducible. Such information would enable AIDS investigators to develop an assortment of standardized assays that could be offered in the form of digitalized assay scripts. Second, by offering common resources to participating investigators (e.g., cell cultures, growth media, viral stocks), batch science facilities would help to eliminate many of the unknowns that limit comparability. The high-throughput environment would permit enormous numbers of experiments to be run in parallel and against a series of standards on a routine basis to maintain reproducibility and reliability.
INTELLECTUAL PROPERTY
With high-throughput technologies changing the means by which research is conducted, it will be important to maintain the traditional reward system for AIDS investigators. At the heart of this system is the freedom to decide how to share data and new information, which can lead to scientific publications and credit for discoveries involving intellectual property (National Research Council, 1997). For each category shown in Box 6.4, data ownership and privileges can be assigned according to the source of financial support.
For the closed category, data would belong solely to the commercial organization (such as a pharmaceutical company) that submitted samples
|
BOX 6.4 Data Ownership and Privileges
|
and assay scripts and paid for the research or testing services. Upon completing such work, the batch science facility would encrypt and forward all the raw data to the purchasing organization. Afterwards, it would be the organization's responsibility to manage the security of its private property. For a period of time, the facility would also maintain a secure copy of the digital records to assure redundancy and integrity in accordance with contractual agreements.
For the principal investigator category, data would belong to the person receiving government grant support for a reasonable period of time—for example, for as long as 2 to 3 years after the grant ends. Good digital practices would be tied to ongoing grant support, requiring each investigator to maintain his or her database records in an orderly manner. After the time embargo had expired, relational links would be attached to the investigator's digital records and the information would become available to others.
For the consortia category, data would belong to all of the collaborating investigators for a reasonable period of time, as suggested above. The collaborators would also have responsibility for maintaining their digital records in an orderly manner, most likely under the supervision of the
group's database manager. After the time embargo expires, the organized information would become available to others.
For the open category, data and its associated links would belong to the public after its quality is assured. The digital records would come from voluntary submissions and time embargoed data that would be released automatically. The main issues would be maintaining backup copies to assure integrity and deciding how to inventory the data and build relational links.
The National Center for Biotechnology Information (www.ncbi.nlm.nih.gov), the HIV Sequence Database (www.hiv.lanl.gov), the European Molecular Biology Laboratory (www.embl-heidelberg.de), and the DNA Data Bank of Japan (www.ddbj.nig.ac.jp) are examples of public database facilities in North America, Europe, and Asia, respectively. Some of these organizations have already established guidelines regarding time-embargoed ownership of data, which could provide reasonable starting points for standard agreements pertaining to batch science (Layne and Beugelsdijk, 1998).
IMPLEMENTATION
A blueprint for initiating batch science via the Internet would involve four related areas of effort. Hardware integration would deal with building particular batch science machines from commercial hardware, converting commercial modules to SLM-based operation, and constructing machines that work together to serve investigators' needs. Software integration would deal with creating a flexible suite of SLM controllers, task sequence controllers, and process control tools that can be used for a variety of AIDS research efforts. Adapting science would deal with quality control matters, standardization of assays, and the scaling-up of necessary reagents so that automated instruments can reach their greatest potential. Laboratory containment would deal with the design, operation, and maintenance of bio-hazard facilities dedicated to high-throughput infectious disease research. Of further significance, batch science facilities would reduce occupational exposures to infectious agents and reduce the unit cost of laboratory experiments by taking advantage of economies of scale (see Box 6.5).
AIDS investigators working at global pharmaceutical companies have had strategic access to high-throughput automated laboratory instruments, while investigators working at most major universities have enjoyed far less access to such experimental facilities. This skewed situation, however, is ripe for change. To maintain a pipeline for creating new therapeutics, global pharmaceutical companies are forming increasing numbers of alliances with external academic research groups (Herrling, 1998). Consistent with the trend in research “outsourcing,” batch science facilities
|
BOX 6.5 Batch Science Facilities Batch science facilities would take advantage of economies of scale, thereby reducing the unit cost of laboratory experiments. Below is an economic comparison of a manual laboratory (employing 100 full-time technicians and occupying ~50,000 sq. ft. of floor space) to a batch science facility (employing 10 full-time technicians and occupying ~5,000 sq. ft. of space) with the same throughput capacity. Capital costs. Typical biohazard level 3 (BL3) space costs $200 per square foot, and new equipment costs $200 per square foot. In comparison, high-throughput laboratory space costs $400 per square foot, and automation equipment costs $1,000 per square foot. The cost of human labor scales as a linear relationship, whereas the cost of batch science scales as a sublinear one.
Operating costs. Typical salary and fringe benefits for a laboratory technician cost $50,000 per year. One technician can perform approximately 400 detailed laboratory procedures annually at an average cost of $250 per procedure, which scales linearly. Automated instruments are more mechanically adept than skilled technicians, and the relative savings could easily amount to a fivefold reduction per test ($50 per procedure) when compared to human-based manipulations.
Five-year average. Over 5 years both facilities would perform a total of 200,000 major laboratory procedures. Batch science facilities would reduce the unit cost of laboratory testing by over threefold. The overall savings would make it possible for investigators to carry out larger experimental undertakings for the same dollar expenditures.
|
they would thus serve as national and international centers for carrying out investigator-initiated research, they would be sponsored by consortia of government agencies and research-driven companies, and would be located at universities or national laboratories. During the start-up period, the center would receive major financial support from sponsoring organizations, permitting the creation and validation of its initial resources. After a center becomes fully operational, the level of start-up support would be scaled down and, over several years would feasibly evolve into a nonprofit
entity that charges investigators reasonable fees for “mass-customized” testing services.
All too often, AIDS investigators shy away from problems that deal with the bigger picture simply because they lack the necessary tools to move ahead. In HIV/AIDS research, there are many important problems to be solved that seem beyond reach because of the enormous “experimental spaces” to be explored and characterized (Fields, 1994). The HIV-RNA genome is composed of ~104 bases, which corresponds to ~3,000
amino acids. If only a small fraction (~1 percent) of amino acids were responsible for conferring unique viral properties, and only one specific amino acid (from the total selection of 20) conferred this property, there could be upwards of 230 ≈ 109 unique variations (see Figure 6.4). Even if only one in a thousand mutations produced viable offspring, there would still be ~106 variants, which is an enormous number to attempt to sample and characterize. At present, we have only sketchy information mapping genotypic variations to phenotypic expressions, making it difficult to relate the significance of such variations (if any) to therapies and vaccine development (Figure 6.1). And what little knowledge we do have is hopelessly scattered among various investigators' notebooks. Batch science facilities, with the capacity to perform the work of hundreds of laboratory technicians, would enable investigators to conceptualize and attack problems from entirely new directions. For many such problems with large phase spaces, high-throughput laboratory facilities with digitalized record-keeping are perhaps the only feasible means for moving ahead.
Health care is progressing in the direction of therapies that are precisely tailored to individuals or smaller groups of patients. In this regard, HIV-infected individuals who take antiviral drugs must rely on precisely the “right” combination (e.g., reverse transcriptase plus protease inhibitors) to halt viral replication (Deeks and Abrams, 1997). With current clinical and laboratory practices, however, there is no single test to establish the optimal regimen. Therefore, to maintain effective therapies, physicians have come to rely on a series of tests (i.e., viral loads, molecular sequencing assays, drug susceptibility assays) that are highly repetitive, time consuming, and labor intensive. Batch science facilities would assist by providing the necessary high-throughput testing services and creating an organized base of information for clinical decision making. In addition, pharmaceutical companies could use batch science facilities to track the molecular evolution of HIV in patients around the world. A wider base of information could then be used to guide the development of new generations of antiviral drugs.
In the Third World, dedicated AIDS investigators are often forced to work with primitive laboratory resources. For such infectious disease research, however, this is precisely where any number of meaningful studies should be conducted. With simple resources in the field—an Internet connection, plasticware, barcodes, refrigeration, and air transportation—one batch science facility could serve large geographical regions.
The creation of batch science facilities will not require a Manhattan Project (nor major redirections of resources) as some might suspect. On the contrary, it will involve a focused effort by a relatively small team of motivated engineers and scientists. With appropriate support, the first-generation facilities could be running within 2 to 3 years, and next-
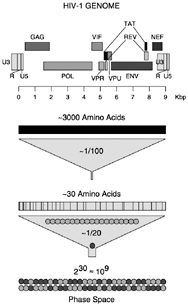
FIGURE 6.4 Estimating the number of HIV-1 isolates by combinatoric arguments. The entire HIV-RNA genome codes for ~3,000 amino acids and mutations in many of these positions may lead to viable viruses. If we conservatively assume that only ~30 of these positions (~1/100) are responsible for conferring phenotypic properties and that only one amino acid out of a pool of 20 (~1/20) is capable of conferring viable viruses, there could be upward of 230 ≈ 109 viable genotypes and unique phenotypes. There are many ways to pose such combinatoric arguments, but they all generate large numbers.
generation facilities would soon follow. If we assume that the facilities perform the work of, say 100 laboratory technicians at a fraction of the cost, the development costs would be recouped within several years. The time is right for government funding agencies, the World Health Organization, health-related foundations, and global pharmaceutical companies to consider their respective roles in building “intermediate-scale” research infrastructures that are comparable to those in the Human Genome Project. AIDS investigators working in basic science, clinical trials, and public health efforts will have no problems in finding clever (and unforeseen) ways to use flexible laboratory firepower.
REFERENCES
American Society for Testing and Materials (ASTM), Subcommittee E-49.52 1998. Laboratory Equipment Control Interface Specification (www.thermal.esa.lanl.gov/astm).
Anderson, R. M., C. A. Donnelly, and S. Gupta. 1997. Vaccine design, evaluation, and community-based use for antigenically variable infectious agents. Lancet, 350:1466-1470.
Baggiolini, M. 1998. Chemokines and leukocyte traffic. Nature, 392:565-568.
Bangham, C. R. M., and R. E. Phillips. 1997. What is required of an HIV vaccine? Lancet, 350:1617-1621.
Beugelsdijk, T. J. 1989. Trends for the laboratory of tomorrow. Journal of Laboratory Robotics Automation, 1:11-15.
Burton, D. R. 1997. A vaccine for HIV type 1: The antibody prospecting. Proceedings of the National Academy of Sciences USA, 94:10018-10023.
Callan, M. F., L. Tan, N. Annels, G. S. Ogg, J. D. Wilson, C. A. O'Callaghan, N. Steven, A. J. McMichael, and A. B. Rickinson. 1998. Direct visualization of antigen-specific CD8+ T cells during the primary immune response to Epstein-Barr virus in vivo. Journal of Experimental Medicine, 187:1395-1402.
Chumbley, L. S., M. Meyer, K. Fredrickson, and F. Laabs. 1995. Computer networked scanning electron microscope for teaching, research, and industrial applications. Microscopy Research Technique, 32:330-336.
Deeks, S. G., and D. I. Abrams. 1997. Genotypic-resistance arrays and anti-retroviral therapy. Lancet, 349:1489-1490.
Fields, B. N. 1994. AIDS: Time to turn to basic science. Nature, 369:95-96.
Garner, H. R. 1997. Custom hardware and software for genome center operations: From robotic control to databases. Pp. 3-41in Automation Technologies for Genome Characterization, T. J. Beugelsdijk, ed. New York: John Wiley.
Gazzard, B. G., and G. J. Moyle, on behalf of the BHIVA Guidelines Writing Committee. 1998. 1998 revision to the British HIV Association guidelines for antiretroviral treatment of HIV seropositive individuals. Lancet, 352:314-316.
Germain, E. 1996. Fast lanes on the Internet. Science, 273:585-588.
Hecht, F. M., R. M. Grant, C. J. Petropoulos, B. Dillon, M. A. Chesney, H. Tian, N. S. Hellmann, N. I. Bandrapalli, L. Digilio, B. Branson, J. O. Kahn, et al. 1998. Sexual transmission of an HIV-1 variant resistant to multiple reverse-transcriptase and protease inhibitors. New England Journal of Medicine, 339:307-311.
Heilman, C. A., and D. Baltimore. 1998. HIV vaccines—Where are we going? Nature Medicine (Suppl.), 4:532-534.
Herrling, P. L. 1998. Maximizing pharmaceutical research by collaboration. Nature (Suppl.), 392:32-35.
Hirsch, M. S., B. Conway, R. T. D'Aquila, V. A. Johnson, F. Brun-Vezinet, B. Clotet, L. M. Demeter, S. M. Hammer, D. M. Jacobsen, D. R. Kuritzkes, C. Loveday, J. W. Mellors, S. Vella, and D. D. Richman. 1998. Antiretroviral drug resistance testing in adults with HIV infection: Implications for clinical management. International AIDS Society—USA Panel. Journal of the American Medical Association, 279:1984-1991.
Lanier, B. M., V. G. Cerf, and D. D. Clark. 1997. The past and future history of the Internet. Communications ACM, 40:102-108.
Layne, S. P., and T. J. Beugelsdijk. 1998. Laboratory firepower for infectious disease research. Nature Biotechnology, 16:825-829.
Layne, S. P., M. J. Merges, M. Dembo, J. L. Spouge, S. R. Conley, J. P. Moore, J. L. Raina, H. Renz, H. R. Gelderblom, and P. L. Nara. 1992. Factors underlying spontaneous inactivation and susceptibility to neutralization of human immunodeficiency virus. Virology, 189:695-714.
Martinez-Picado, J., and R. D'Aquila. 1998. HIV-1 drug resistance assays in clinical management. AIDS Clinical Care, 10:81-88.
Moxon, R. E. 1997. Applications of molecular microbiology to vaccinology. Lancet, 350:1240-1244.
Murray, C. J. L., and A. D. Lopez. 1997. Alternative projections of mortality and disability by cause 1990-2020: Global Burden of Disease Study. Lancet, 349:1498-1504.
National Research Council, Computer Science and Telecommunications Board. 1993. National Collaboratories. Washington, D.C.: National Academy Press.
National Research Council, Committee on Intellectual Property Rights and Research Tools in Molecular Biology. 1997. Intellectual Property Rights and Research Tools in Molecular Biology Washington, D.C.: National Academy Press.
Pantaleo, G., H. Soudeyns, J. F. Demarest, M. Vaccarezza, C. Graziosi, S. Paolucci, M. Daucher, O. J. Cohen, F. Denis, W. E. Biddison, R. P. Sekaly, and A. S. Fauci. 1997. Evidence for rapid disappearance of initially expanded HIV-specific CD8+ cell clones during primary HIV infection. Proceedings of the National Academy of Sciences USA, 94:9848-9853.
Persidis, A. 1998. High-throughput screening. Nature Biotechnology, 16:488-489.
Pollard, M. J. 1997. Automation strategies: A modular approach. Pp. 43-63 in Automation Technologies for Genome Characterization, T. J. Beugelsdijk, ed. New York: John Wiley.
Rosenberg, E. S., J. M. Billingsley, A. M. Caliendo, S. L. Boswell, P. E. Sax, S. A. Kalams, and B. D. Walker. 1997. Vigorous HIV-1–specific CD4+ T cell responses associated with control of viremia. Science, 278:1447-1450.
Schwartz, R. S. 1998. Direct visualization of antigen-specific cytotoxic T cells—A new insight into immune disease. New England Journal of Medicine, 339:1076-1078.
Spouge, J. L., and S. P. Layne. 1999. A practical method for simultaneously determining the effective burst sizes and cycle times of viruses. Proceedings of the National Academy of Sciences USA, 96:7017-7022.