Mendel in the Kitchen: A Scientist's View of Genetically Modified Foods (2004)
Chapter: 5 Tinkering with Evolution
5
TINKERING WITH EVOLUTION
What distinguishes a butterfly from a lion, a hen from a fly, or a worm from a whale is much less a difference in chemical constituents than in the organization and the distribution of these constituents.
—François Jacob (1977)
By 1804 Julius von Sachs knew that the carbon and nitrogen in a plant were precisely the same as those in a rock: they were not imbued with a vital force just because they resided in a living thing. In 2001 Craig Venter and the other scientists who sequenced the human genome and compared it to the genomes of yeast and fruit flies and mice and maize and the little weed Arabidopsis concluded that genes, too, were not “the mechanisms that generate the complexities” of a creature.
Humans have roughly 200 types of cells: blood cells, bone cells, skin cells, liver cells, heart cells, and more. All come from a single cell, the fertilized egg, over a period of years. Yet humans have only 10 times as many genes as the bacterium E. coli, itself a single cell. We have twice as many genes as a fruit fly. But a corn plant has a genome as large as ours. Our complement of genes is startlingly similar to that of mice. Set the human sequence and that of the mouse side by side, and only some 300 human genes will have no counterpart in the mouse. Anthropologist Svante Pääbo wrote in 2001 that such cross-species
comparisons “make the unity of life more obvious to everyone.” They are “a source of humility and a blow to the idea of human uniqueness.”
In 1977, well before the first genome was parsed into its A’s, T’s, G’s, and C’s, François Jacob had raised that very question in an essay called “Evolution and Tinkering”: “What distinguishes a butterfly from a lion, a hen from a fly, or a worm from a whale?” Answering himself, Jacob, a molecular geneticist, made the radical suggestion that evolution was more like tinkering than engineering. Each organism’s adaptation to its niche is far from perfect—this fact Darwin himself had been keenly aware of. Yet in the years after The Origin of Species, the concept took hold that evolution worked like an engineer designing a machine. Imperceptibly, natural selection was endowed with the ability to optimize. The idea that every bit of a worm or a whale was there for a reason—one understandable through the filter of natural selection—is still cherished by many, if not most, biologists today.
Jacob was saying something quite different. An engineer starts with a plan, and with the materials specified, then designs the product with the appropriate methods to be as perfect as possible. A tinkerer starts with whatever is at hand and makes something that does the job, even if far from perfect. A tinkerer needs no plan—just a supply of spare parts. These, evolution provides in abundance. However life on Earth arose, most likely it did not start twice. Thus each evolutionary change occurred in an organism that already existed. Those organisms, for most of life’s history on Earth, were single cells. (Most living things, at least in numbers, still are.)
Today’s many creatures show indisputable differences: Hens have no teeth, worms have no feathers, E. coli has no flowers, potatoes have no toes. Yet much is the same. Evolution seems to copy a part, then change it a bit, tinkering with it to add a new function.
A cell, whether of E. coli, a lion, a fly, or a lettuce, is a jelly-like agglomeration of long molecules, most of which are proteins. The cell is encased in a membrane made of fats, and threaded through with other membranes. At its center is DNA. In higher organisms (from yeast to people) the DNA, in its spindly chromosomes, is inside a separate compartment, the nucleus. In lower organisms (bacteria) the
single chromosome is tightly bundled, but not segregated from the rest of the cell.
The cells of higher organisms also house mitochondria. These small structures are descended from bacteria that took up residence inside other cells long ago. Mitochondria have their own DNA, although many of the original bacterial genes have moved into the cell’s chromosomes. Inside the mitochrondria are proteins that break down sugar, providing energy for the cell to move and grow. The cells of green plants have an additional kind of cell-within-a-cell, or organelle, the chloroplast. Chloroplasts let plants extract energy from light—a spectacular ability. Chloroplasts are descended from cyanobacteria, a kind of bacteria that learned to harvest light energy more than a billion years ago. They too still have their own DNA. But, just as in mitochondria, many of their original genes have migrated into the nucleus and now reside on the cell’s chromosomes.
There are variations on the theme of “cell.” Blood cells in mammals fill up with iron-rich hemoglobin, then eliminate their nuclei to focus on toting oxygen from the lungs to every cell. Plants develop stacks of long cells, end to end. These interconnect, then die, forming
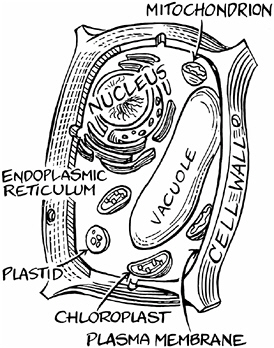
The interior of a plant cell
long straws that pull water out of the ground. But such specializations came late in evolution. From the simplest organism to the most complex, the cells of all living things have much in common. Every time a cell divides, for example, all its DNA must be copied. The copying machine in the bacterium E. coli is made up of 10 different proteins: some form the structure of the machine, some latch it onto the DNA, and some link the letters ATGC at a rate of 500 per second. In 20 minutes the copying is done. To copy a human cell’s much longer DNA takes about eight hours, with a thousand similar machines.
The process of making a protein is also not much changed from E. coli to Homo sapiens. Each protein is coded for by a gene, a length of DNA that can be hundreds or thousands (or even hundreds of thousands) of letters long. The gene is first transcribed into RNA (ribonucleic acid), like a television interview transcribed for print. Then the transcript is translated from RNA into protein, as if from Japanese to English. RNA is similar in structure to DNA (deoxyribonucleic acid), except that it contains a different sugar (ribose instead of deoxyribose) and it has only one strand, not two, so it doesn’t form the familiar double helix. Like DNA, RNA is made of four building blocks, called bases or nucleotides. Both molecules have the bases A (adenine), G (guanine), and C (cytosine). RNA, however, has U (uracil) in place of the T (thymine) in DNA. The transcript is a long molecule of RNA called messenger RNA. The translation into protein is the job of a number of smaller forms of RNA called transfer RNAs.
All of each gene’s DNA—both the protein-coding exons and the mysterious introns—is read and transcribed into an RNA molecule. The chromosome itself is part of this process. Chromatin, the complex of DNA and proteins from which chromosomes are made, has to be remodeled to allow transcription to occur. A large transcription machine must be assembled. This machine unzips the DNA double helix at the appropriate spot and allows the RNA letters room to bind with their partners on one of the DNA strands. As soon as the transcript begins, the leading end is linked to a protein. This protein attracts others, whose job it is to detect the beginning and end of each intron. They cut the transcript there and splice the two exon ends together, kicking out the intron.

From DNA to protein
When the transcription machine reads a certain signal in the DNA, it stops. A new protein machine then adds a long string of A’s to the transcript, familiarly called a “polyA” tail. The tail is the transcript’s certificate of accuracy. Throughout the transcription process, layers of proofreaders (proteins again) have been checking for mistakes. The tail is the last rubber stamp. Only when it has a polyA tail can a transcript travel out of the nucleus and be translated into a protein.
In bacteria these two processes—transcription and translation—are completely coupled. Because bacteria are simpler and have fewer (and simpler) genes, they seem to be able to get away without proof-reading. In higher cells the transcript, the certified error-free RNA message, goes out of the nucleus and into the cytoplasm—the bulk of the rest of the cell. There the message is read by a ribosome, another machine made of many proteins. The ribosome follows a strict rule of
translation: each set of three nucleotides, each codon, in the messenger RNA is recognized by a matching anticodon at one end of a molecule of transfer RNA. The messenger RNA’s UUU, for instance, is recognized by the transfer RNA’s AAA.
On its tail end, each transfer RNA has attached to it a particular one of the 20 amino acids, in this case the amino acid phenylalanine. The transfer RNAs take their places along the messenger RNA, lining up the amino acids on their tails in the order specified by the DNA code. The amino acids are linked in a chain and then released from the transfer RNA. The long chain of amino acids then folds up to make a functioning protein. Each kind of protein folds, like yarn into a ball, in its own way, although there are common themes, called “folds,” that proteins favor. The sequence of amino acids determines the shape of the protein. The shape of the protein determines what it does.
Curiously, certain protein folds have been especially successful in evolution—all of the hundreds of thousands of proteins fold into no more than a few thousand of these overall structures. And many proteins have proved so useful that their sequences have changed very slowly over the eons. These similarities mean that when a new protein is identified, scientists routinely check its sequence and shape against the databases of those already known. They can often guess what the new protein does from the functions of others that it resembles.
Proteins actually do a surprisingly small number of jobs. They make and break chemical bonds. They bind and transport small molecules, such as hormones, sugars, or oxygen. They bind to nucleic acids (DNA and RNA), to membranes, and to other proteins, giving structure to cells and regulating their chemical activities. Each of these jobs is done by different protein parts, called domains. Protein domains often come in pairs, so a particular protein might bind to DNA with one domain and break a chemical bond in that DNA with another domain. Groups of three or more domains that work together are also common, increasing the variety of jobs a single protein can do. Protein domains are the spare parts of evolutionary tinkering: different proteins have different combinations. By shuffling domains or adding new ones, an old protein can acquire a new function. Yet if you compare a domain from a protein in a bacterium, a plant, and a person, they might look quite similar—both in sequence and in structure.
The similarity among proteins in different organisms reflects not only the underlying fact that the cells of all organisms carry out many of the same chemical reactions, but also the evolutionary relationships among the organisms. Similarity at the protein level doesn’t mean that all living creatures are the same—any more than a Model A Ford is the same as an SUV. It means that they’re made of parts that do the same kinds of tasks, and it means that organisms evolved gradually. It also doesn’t mean that every part in an SUV looks or works the same as its counterpart in the Model A. But because the rules for transcribing genes and translating proteins are the same in all living things, a gene from a bacterium can, with the proper switches added, work in a plant cell. The plant cell will make the very same protein from that gene that the bacterium made.
![]()
In the early 1990s Pam Dunsmuir and her colleagues at DNA Plant Technology Corporation, a biotechnology company in California, put a fish gene into a tomato. The experiment has since become a rallying point for protests against genetically modified foods. A Canadian writer, Jennie Addario, reported on one such event at a Loblaws grocery store in Toronto in 2002:
Suddenly, about 50 protesters march toward the grocery store, chanting, “Hey hey, ho ho, leave our DNA alone.” One demonstrator beats a drum. Others hold up homemade placards urging shoppers to “Be GMO Free” and warning them that “You’re Eating Genetically Modified Organisms.” Some protesters even wear costumes. One young woman sports a hand-drawn cardboard ear of corn that hangs from her neck like an over-sized tie. Another is far more creative: she is dressed from top to bottom as a gigantic, plump, red tomato, complete with a large stem and leaf jutting from her head and, bursting out from her vegetable belly, a fish head with eyes, fins, and gills. So it was no surprise that in the next day’s edition of The Toronto Star a picture of the mutant tomato-fish received prominent play. In the story accompanying the photograph, the reporter, Tanya Talaga, wrote that the protester “said her costume represented the fact that fish genes are spliced into tomatoes to give them a longer shelf life.” The impression given to the reader by this statement and the rest of Talaga’s story was that all the tomatoes in grocery stores, including those at Loblaws, were swimming in fish genes. But that is simply not the case. In fact, there are no supermarket tomatoes anywhere that are packed with fish genes.
The “mutant tomato-fish” is nothing more than an urban legend, writes scientist Alan McHughen in his book, Pandora’s Picnic Basket: The Potential and Hazards of Genetically Modified Foods. It is a story made of “shadows” to “incite public fear by creating an unnecessary scare.”
Dunsmuir’s experiment did not show the effect she was looking for, and a tomato containing a fish gene was never grown or commercialized. Nor did increasing shelf life have anything to do with it—that was another tomato, Calgene’s Flavr Savr, which was the first genetically engineered whole food to come to market. First sold in 1993, the Flavr Savr contained no fish genes. In fact, it contained no foreign genes at all. The modified gene was a normal tomato gene that coded for an enzyme, polygalacturonase (PG), involved in the fruit ripening process. The researchers had copied the PG gene, flipped it, and reinserted it in reverse order. “The resulting ‘antisense’ PG gene, by some unknown mechanism,” wrote one of the researchers, Belinda Martineau, in her book First Fruit: The Creation of the Flavr Savr Tomato and the Birth of Genetically Engineered Food, “shut down native PG protein production in the engineered plants.” Ten years later we know that the “shutdown” is a normal plant response that protects from invasion by viruses.
According to Martineau, the Flavr Savr “sold like hotcakes.” “Bert Gee, the owner of State Market, resorted to limiting customers to two Flavr Savr tomatoes a day,” Martineau writes. He sold “gift packs—‘four tomatoes in a box to give to friends and family.’” A waiting list formed of grocers wanting to stock Calgene’s product. But Calgene couldn’t deliver. Belatedly they discovered that they “had to handle vine-ripened Flavr Savr tomato varieties, it seemed, just as gently as any other, conventionally developed vine-ripened fruit.” The assumption that a tomato slow to rot would stay firm enough to pack and ship “wasn’t panning out.” Their $2-per-pound tomatoes cost the company $10-per-pound to deliver. Flavr Savr was off the market by 1996. Calgene, near bankruptcy, was sold to Monsanto.
The tomato with the fish gene, on the other hand, never made it out of the lab. Although the gene itself worked properly—it produced the same protein in the tomato as it did in the fish—the protein did not have the desired effect, which was to protect the tomato plant from frost in the field and the tomatoes from cold damage on the way to
market or at home in the refrigerator. The gene came from a type of Arctic flounder; the protein it encodes keeps the fish from freezing in icy waters. When cells freeze, the transition from water to ice breaks them apart just as surely as leaving a bottle of beer in the freezer breaks the bottle. Yet many organisms, both animals and plants, avoid such severe freeze damage. Seeds, all trees, and many bushes do, as well as salamanders, snakes, insects, and some fish. Just as we use salt to de-ice airplanes or roads, they use high concentrations of salt or other substances that keep water from freezing until the temperature is colder than normal.
However, 30 years ago scientists working in the Antarctic found fish that fought freezing a different way. Special proteins in their blood allow them to survive temperatures that would burst even salt-rich cells. These antifreeze proteins lower the freezing point through a mechanism that, unlike the salt method, does not depend on their concentration. As the ice crystals are forming, the proteins bind to the prism faces of the ice. The ice crystals grow more slowly and so do not disrupt the fish’s cells.
Such antifreeze proteins have since been found in flatfishes, sculpins, sea ravens, smelt, herring, eel pouts, northern cods, and wolffishes. They are found in mealworms and spruce budworms, and in the larvae of the fire-colored beetle. Winter rye has them, as do carrots and bittersweet nightshade, a plant in the same genus as tomato, Solanum. All of these proteins work in the same basic way, although how they are related to one another is not clear. One of the fish proteins appears to have evolved from a protease, an enzyme that breaks down proteins. Others, isolated from winter rye, originated from chitinase genes recently enough that some of them still retain their earlier function, which is to break down polysaccharides, a kind of sugar polymer.
The antifreeze proteins in winter rye and carrots and nightshade are not as effective as those in fish or insects, and so scientists have continued their efforts to insert new antifreeze genes into plants. The goal of most such experiments is to protect crops better from untimely frost. A gene from the fire-colored beetle, for instance, was recently engineered into Arabidopsis thaliana, the laboratory plant now popu-
lar with plant geneticists. The insect protein worked quite well, lowering the temperature at which the plants froze.
Why did it work? First, of course, the rules for transcribing and translating DNA into proteins are everywhere the same. The DNA codon AAA always stands for the amino acid phenylalanine; CTT always means glutamine. The amino acid sequence is the same whether the gene resides in a fish or a beetle or Arabidopsis or a tomato. Among organisms like these, whose evolutionary paths diverged so long ago, there are of course subtle differences. Each has its own way of saying when and where a gene will be expressed. They differ in how often they use a certain codon—with 64 possible triplets but only 20 amino acids, two or more codons can be synonyms. But the triplet code itself is universal. A gene directs the construction of precisely the same protein in whatever organism it finds itself in. The second reason the new protein works is that a protein’s task, always and everywhere, is determined by its sequence and how it folds.
The fact that many different kinds of proteins evolved to prevent freezing in many different organisms illustrates just the kind of tinkering Jacob had in mind. Some of these proteins are efficient at preventing freezing, some less so. But they are all simply proteins. It is as impossible to attach the quality of being an insect to a protein as it is to attach the quality of being a car to a spark plug. There is no way a gene from a fish could make a tomato “fishy.” When the fire-colored beetle gene is expressed in Arabidopsis, the plant is still Arabidopsis. Only now it can express one extra protein, one that changes how ice crystals form.
Whether the antifreeze protein will do for Arabidopsis (or a tomato) what it did for the beetle (or the fish) from which the gene was taken is more difficult to predict. It has to be tested, as Dunsmuir was doing. Each organism modifies its proteins a little differently. These modifications can determine the fate of the protein within the cell, as well as what partners the protein works with. As organisms become more complex, their proteins become more and more exclusive. They are pickier about where they carry out their functions and whom they partner with. They will work only in the nucleus or only in the mitochondrion or only on the cell’s outer surface. Or the gene that encodes a protein will be expressed in one cell type, but not another.
For example, hemoglobin binds oxygen in both animals and plants. In animals, hemoglobin proteins are made in red blood cells that circulate in the blood stream to bring oxygen to cells far from the lungs. In legumes, like peas and beans, they are used for a different purpose: to create the perfect environment for the nitrogen-fixing bacteria in the plant’s roots. The enzyme that converts nitrogen from the air into compounds the plant can use to grow (so that it does not need nitrogen fertilizer) can’t function in the presence of oxygen. So the plant builds an oxygen-free home for this beneficial bacterium, lining special root nodules in which the bacteria live with hemoglobin molecules to keep oxygen away. The plant and animal hemoglobin molecules are not identical—they’ve drifted apart over evolutionary time. But their ability to bind oxygen is much the same and their similarity in structure is still recognizable.
So to paraphrase François Jacob, the difference between a person and a pea is less a difference in their genes than in how, where, and when those genes function and the proteins are deployed. The difference between a fish and a tomato is not so much in what their proteins do, but in how much of each is made, where it is made, and how it interacts with its partners.
With this new understanding of proteins comes a deeper understanding of evolution. Evolution once seemed quite linear—and human beings were obviously as near to perfection as any organism had come. But some complex animals, such as horseshoe crabs and coelacanths, have persisted unchanged for eons. We call them, quaintly, living fossils—as if they ought to be extinct. Indeed, many species of complex animals are extinct, even as simpler organisms have persisted. Modern bacteria are not very different from fossil bacteria. They have the same basic single-celled layout, despite many specializations and diversifications. Newer life forms do not replace older ones because they are more complex, higher, or better adapted. Instead, the evolution of life on Earth is marked both by increasing complexity and by boisterous diversity. The diversity is easier to understand than the complexity. Many organisms can be derived from an original type with just a little bit of tinkering. From ordinary fish, living deep in caves where light never penetrates, come blind fish without eyes.
But where does the complexity come from? If evolution begins with an existing fish and then adapts it, how can something essential ever change? How can a gene be rearranged to give rise to a new protein with a new function or a new pattern of expression if the original protein is one the organism can’t live without?
One answer is to have spare copies of genes. Often a single gene is copied—then copied again and again, with all of the duplicates staying in place, side by side. Sometimes a whole section of chromosome is doubled—or even the whole genome. To copy a whole genome is in fact quite simple. The machinery is already in place, because every time a cell divides, all of its DNA—the whole genome—is copied. Sometimes both copies go into one cell by mistake when the cell splits into two daughter cells. Occasionally, the daughter cell with a doubled genome is a germ cell—a sex cell—and so the doubled genes are handed down to the next generation. This, incidentally, is how colchicine works, the chemical from the autumn crocus with which plant breeders created triticale and the seedless watermelon. When the genome is copied, colchicine keeps both sets together.
Extra copies of genes, chromosomes, or whole genomes are made in the simplest organisms and in the most complex. But the copies tend to persist in eukaryotes, organisms whose cells have nuclei, much more often than they do in prokaryotes, the bacteria. Bacteria have a handful of duplicated genes; humans, mice, and corn plants (all eukaryotes) have many thousands.
Just why extra copies are made and why they persist, swelling genomes, is not well understood. But there is no doubt that genomes grow in the course of evolution by duplicating their DNA. Once a gene is doubled, changing the protein is easy: one copy of the gene supplies the original protein; the other copy can change. A duplicated gene can be lost—and they often are—or it can make itself essential by acquiring a new function—and they occasionally do. Many, if not most, genes in plants and animals belong to clusters, or families, of related genes. Some gene families are small, with as few as two members; others are huge, with hundreds or even thousands. Each family arose by copying and tinkering with a previous DNA sequence.
![]()
If copying is the key to complexity it should be no surprise that the genomes of animals and plants are full of transposons, or jumping genes. But when Barbara McClintock announced in 1948 that genes could move from one place to another, it was tantamount to saying the kitchen could occasionally move into the attic. Geneticists—McClintock among them—had been mapping genes for decades. They knew very well that genes maintained their locations—maps couldn’t be made if genes moved around all the time. Genes had fixed locations, like beads on a string, and their order along the chromosomes could be counted on generation after generation. Occasionally chromosomes exchanged parts—this was called a translocation—but these were unusual events. Genes didn’t jump. Yet McClintock reported that they did.
McClintock began her work at Cornell University in the early 1920s, at the same time as George Beadle was beginning his studies there of corn (maize) and teosinte. McClintock was curious about the relationship between chromosomes and the inheritance of traits. To understand it, she too worked with corn.
From the account of one of McClintock’s biographers, you would think that the genetics of corn was a gentlewomanly pursuit: she “watches the plants grow over the summer, and spends the long quiet winters analyzing the results.” A scientist who worked with McClintock reported a very different experience: “Doing maize genetics is hot and sweaty work—sunup to sundown—sometimes in temperatures over 100 degrees. And then there’s keeping track of what crosses have been done and what crosses must be done the next day. Every plant and every cross must be recorded before the next day’s crosses are planned.” She remembers being so exhausted by the end of two months of 5:00 A.M. to 2:00 A.M. days that she seriously considered abandoning the project. Even McClintock admitted that it took her months to recover from each summer’s work.
When corn flowers, pollen bursts from the tassels at the plants’ tops. Millions of grains swirl through the air, to be snagged by the long silks protruding from the female flowers, the ears. Controlling this boisterous mating process takes extreme care. Each plant is given a
number. Each ear is covered by a bag. When an ear is ready to be fertilized, the bag is briefly lifted and pollen from the desired male is brushed onto the silks of the ear. The tracking numbers of the plants used as the male and the female in the cross are written on the bag. (In today’s genetics labs, these numbers are in the form of barcodes.) Hundreds and hundreds of crosses are done every day at the height of the season. Thousands are done in all. A month or so later, each mature ear is tagged and collected. Then the serious work begins.
A way to visualize the chromosomes of maize had been McClintock’s first major contribution to science. Corn’s chromosomes had been counted—there were 10—but in the 1920s no one could consistently tell one from the next. While a graduate student, McClintock found that the chromosomes in male reproductive cells were the best ones to examine, because they were long and rather stretched out. She adapted a staining technique that had just been developed using a red dye called acetocarmine. The stain made the structure of each chromosome clear. McClintock gave each one a number, distinguishing them by size, by where they attached to each other, and by the pattern of various knobs each one bore. She received her Ph.D. in 1927 and stayed on at Cornell. There she and a student, Harriet Creighton, soon succeeded in connecting the behavior of genes with the behavior of chromosomes. Their paper, published in 1931, has been called one of the “classic papers in genetics,” a “landmark in experimental genetics,” even a “cornerstone.” Its results “are so strong and obvious that their validity cannot be denied.”
This paper made McClintock famous. In 1936 Lewis Stadler, who was then carrying out some of the first mutation breeding experiments on barley, persuaded the University of Missouri to offer her an assistant professorship. There she began to study the behavior of maize chromosomes that had been broken by X-rays. When George Beadle, in the 1940s, was exposing the orange breadmold Neurospora crassa to X-rays—work that would ultimately lead to his Nobel Prize-winning “one-gene, one-enzyme” hypothesis—he asked McClintock to identify the mold’s chromosomes. Much later, in the 1950s, the National Academy of Sciences became concerned that the introduction of high-yielding hybrid corn to Central and South America would result in the
disappearance of local varieties, or landraces, a concern that has resurfaced today in a different guise. The Academy asked McClintock to visit field stations to teach local biologists how to classify types of maize by looking at their chromosomes. She was considered the world’s foremost expert on the structure of the chromosomes of corn.
But the work that most deeply engaged her for more than 50 years, and for which she finally received the Nobel Prize in 1983, at first drove her into obscurity. It began with broken chromosomes. McClintock devised a method for bringing broken chromosomes together from each parent. She saw that the broken ends stuck together, forming a chromosome with two centers—a dicentric chromosome. Normally when a cell divides, it finds a chromosome’s centromere, a special structure at its center, and pulls the chromosome apart at that point. A dicentric chromosome has two centromeres. When the cell divides, it is caught in a tug-of-war. It breaks as the cell attempts to distribute the two centromeres to the two daughter cells. Because it breaks at random, one daughter cell sometimes receives two copies of a chromosome segment (and the genes it carries) and sometimes none.
McClintock saw that the offspring of such plants suddenly showed many new mutations. A startling number of them were the kind called unstable mutations, in which the mutant gene often reverts to its normal function while the plant is growing. These kinds of mutations cause the stripes or spots in decorative Indian corn. Such variegation is also familiar—and highly prized—in flowers like snapdragons and roses.
McClintock found that one mutation seemed to cause a chromosome to break, not at random, as she was accustomed to, but at the same place each time. She began to study this curious gene, which she called Dissociation. Her next discovery was truly momentous: Dissociation could move. In the margins of an unpublished paper on her first Dissociation work are these comments in McClintock’s hand: “At the time, I did not know that Dissociation could change its location. Realization of this did not enter my consciousness until late this spring, following the harvest of the greenhouse crop.” The spring was that of 1948 and the greenhouse crop was grown in the winter of 1947.
Long before McClintock began her work, Rollins Emerson, George Beadle’s advisor, had studied the unstable mutations that cause the col-

The effect of a transposon, or jumping gene, on the colors of a corn kernel
orful patterns in Indian corn. Previous geneticists had simply thrown up their hands when confronted with this peculiar behavior; some dismissed them as “sick” genes from which nothing could be learned. But using a corn variety whose kernels were striped dark red, Emerson found that the genes’ behavior wasn’t so bizarre after all. It could be explained quite well if some kind of inhibitor had gotten stuck to the gene, keeping it temporarily from working. If the gene were needed for colored kernels to be made, the inhibitor would cause colorless stripes. But if the inhibitor fell off now and then while the plant was growing, the gene would work normally: streaks or spots of color would appear. Emerson couldn’t imagine what this inhibitor might be.
Some years later Marcus Rhoades, a lifelong friend of McClintock’s, learned something else about unstable mutations. When he crossed a plant carrying a mutation in a pigment gene (a standard, stable mutation studied by maize geneticists for years) to a variety called Black Mexican Sweet corn, the mutation suddenly became unstable. The instability was caused by a gene he called Dotted. Rhoades’s discovery
meant that a mutation’s stability could depend on the presence or absence of a completely different gene.
McClintock was familiar with all of this work when a new mutation cropped up in her plants in which Dissociation was breaking chromosomes. McClintock noticed an odd-looking kernel. It was largely colorless, but had little colored spots, much like the dots that Rhoades had studied. McClintock knew that Dissociation could move from one place to another in the chromosome. She guessed now that it might occasionally jump into, and disrupt, another gene.
By the time McClintock had tested—and verified—this prediction, she knew the answer to Emerson’s question. The inhibitor was a transposable element (in this case, Dissociation) that could jump into and disrupt the function of a gene. If its activator gene was present (in McClintock’s plants it was called Activator, while in Rhoades’s plants it was Dotted), then the transposable element—today called a transposon—could jump out again.
McClintock’s first major public lecture on transposition was at a Cold Spring Harbor Symposium in 1951. She told an auditorium full of her fellow geneticists that not all genes were fixed in their positions on the chromosomes. Some, like Dissociation and Activator, could jump to new places on the same chromosomes. They could even transpose to a different chromosome. Most of her colleagues were simply puzzled.
By then, McClintock enjoyed an extraordinary reputation as a geneticist. She’d been one of the first women elected to the prestigious National Academy of Sciences. The eminent geneticist Sturtevant, maker of the first gene map, when asked what he thought of McClintock’s work, reportedly replied, “I didn’t understand one word she said, but if she says it is so, it must be so!” Lewis Stadler, who had recruited McClintock to work at the University of Missouri in the 1930s, pronounced it “the most amazing thing.” He added that because of her reputation and great observational skill he had “no choice but to believe her.” Others were less generous, but no less confused.
Because McClintock was on the scientific staff of the Carnegie Institution of Washington’s Department of Genetics, she didn’t have to care what her colleagues thought—or whether the government would fund her research. Her salary was paid. She worked by herself, growing and analyzing corn. She simply did what fascinated her.
Transposons were discovered in the early 1960s in bacteria. In the 1970s and 1980s they were discovered in fruit flies, yeast, worms, fungi, mice—and finally humans. Few organisms lack them. Today we know that genomes are stuffed with transposons and their relatives, called retrotransposons because they multiply outside of chromosomes and then reinsert themselves. Almost half of the human genome and three-quarters of the maize genome consist of nothing but transposons and retrotransposons.
Why did it take so long to discover transposons? Many writers have looked for the answer to this riddle in McClintock’s reclusive character and her supposed inability to communicate. Her first long paper on transposons is compared uncharitably to Watson and Crick’s 800-word note—which one critic described as “tight as a sonnet”—on the structure of DNA. Watson and Crick’s contribution was hailed almost immediately: they were awarded a Nobel Prize a mere nine years later. Three decades would elapse before McClintock received her Nobel Prize.
But the answer doesn’t lie in the person. It is both straightforward and mysterious. The much slower acceptance of McClintock’s discovery, compared with that of Watson and Crick, has a parallel in Mendel’s obscurity and Darwin’s popularity a century earlier. Darwin’s theory of evolution was immediately recognized as important, while Mendel’s laws of heredity weren’t widely integrated into evolutionary thinking for 50 years or more. Darwin’s theory explained things that were easily observed: birds resemble each other in general ways, but each bird is a bit different. It wasn’t at all clear initially what Mendel’s rules of inheritance had to do with the differences that lead to new species.
Equally unimaginable a century later was the importance of jumping genes. Mendel’s laws were by then the central paradigm of genetics. The mapping of genetic loci through the study of mutants was proceeding apace. There was plenty of evidence that genes had fixed locations. Geneticist Ledyard Stebbins’ book The Basis of Individual Variation, published in 1950, acknowledges that there are duplications, inversions, translocations, and deletions in chromosomes. Still, the book reflects the prevailing view that these “are not the materials that selection uses to fashion the diverse kinds of organisms which are the products of evolution.” Instead, he concludes, the majority of evolu-
tionary changes are due to classical genetic point mutations: an A substituted for a G, a T, or a C.
Now another half century has elapsed. In every genome scientists have explored, they find themselves knee-deep in transposons, most of them defective in one way or another—the bones of dead transposons. Those that are not defective are commonly silent, kept inactive by reversible mechanisms. The result is that neither genes nor transposons normally move; chromosome structure is maintained and the transposons are undetectable.
Still, it seems that having an abundance of transposons goes along with being quick at the game of evolution. When the genomes of closely related animals (such as mice and humans) or plants (such as sorghum and maize) are compared, the difference between them lies in the linear relationships among the genes. Often the order is the same for long stretches. Elsewhere, a gene is missing from one genome while persisting in the other. Or a small group of genes occurs in the opposite order in one genome, as if a chromosome segment had been flipped around. Or a gene—or group of genes—has moved from one chromosome to another. Such rearrangements frequently start and end with a transposon. It is hard to escape the conclusion that transposons are central to rearranging chromosomes and moving genes.
Genes can change, they can duplicate and delete, and genomes scramble. It is increasingly evident that what genes do depends more on what they are than where they are—although both a gene’s immediate neighbors and its general genomic neighborhood can influence its expression. But evolution takes a long time—like the movement of tectonic plates. The evolution of a plant is measured in millions of years, not in the months it takes to grow a crop of corn. McClintock saw transposition on this very short timescale only because her experiments with broken chromosomes had awakened sleeping transposons.
It is usually to an organism’s advantage to keep its transposons carefully out of action—silent, asleep. Jumping genes might well power evolution, but they can also do a great deal of damage. When transposons move, they almost always increase in number. The jumps occur just after the chromosome divides, in preparation for cell division. One copy of the transposon stays in its original place, while the
other copy moves to a new location—sometimes on the same chromosome, sometimes on a different one. Each time a transposon moves, it creates a sequence like itself somewhere else in the same genome. When the transposon alights in part of a chromosome that has not yet been replicated, it doubles again, along with the rest of its new chromosome. Then there are three copies of the transposon: one at the original location and two at the new spot.
During the crossing-over phase of cell division, chromosomes exchange parts, or recombine, when the same sequences on two chromosomes line up perfectly with each other. In this way, a new combination of grandmother’s and grandfather’s genes is passed on to the next generation. Normally the chromosomes exchange equal segments, because the same gene is located in the same place on every chromosome. Transposons can disrupt that pattern. Copies at different locations can—and very occasionally do—recombine with each other, causing a translocation, an exchange of parts between different chromosomes. The genes get shuffled. Or a whole segment of the chromosome, bounded by transposons, is duplicated or simply moved to another chromosome.
Transposons often delete portions of themselves as they try to move, damaging and immobilizing themselves. That’s why genomes are full of transposon bits and pieces, often just the very ends. Transposons undergo bursts of activity, then burn out by accumulating defective copies. The Dissociation transposon McClintock first identified was such a defective element, made up of two transposon ends without a middle. Activator, the gene that made Dissociation jump, turned out to be a complete, functional transposon. It supplies Dissociation with the enzyme, called a transposase, that it needs to cut itself out of the DNA and move to a new location.
A transposon that is silent or asleep doesn’t transpose—but neither does it damage itself, so it can survive to become active another day. There are many ways to silence a transposon. A common one in higher plants and animals is one that doesn’t affect the transposon’s genetic structure, so it is called epigenetic—on top of the gene. Small chemical groups, called methyl groups, are added to the DNA. Methyl groups can be put on and taken off without damaging the DNA. But
methylation profoundly affects the way that proteins bind to a gene. The transposase enzymes, for example, that move McClintock’s transposons simply don’t bind to DNA that is covered by methyl groups. So as long as the transposon is methylated it won’t jump.
By bringing together broken chromosomes and initiating cycles of chromosome breakage, McClintock triggered a process that takes off the methyl groups and reactivates the transposons. Years after her initial discoveries, McClintock showed that this chromosome breakage process could wake up Rhoades’s Dotted element too, one she’d never seen active in her plants.
Today we know that the same process can be triggered by irradiating plants with X-rays or gamma rays, or simply through ordinary tissue culture, those methods of regenerating whole plants from clumps of callus cells or from protoplasts. The kinds of crosses that Luther Burbank did between very distantly related plants often have the same results. All of these procedures—wide crosses, tissue culturing, irradiation—have been used extensively in conventional plant breeding. By trial and error, breeders found ways to speed up plant evolution by disturbing and rearranging genomes.